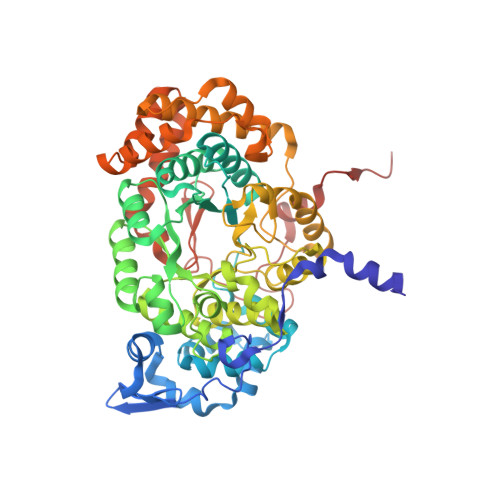
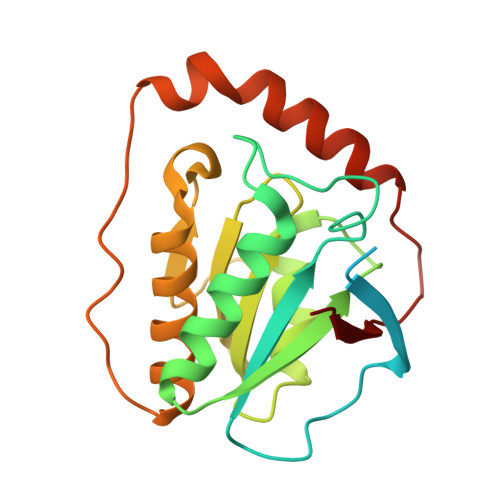
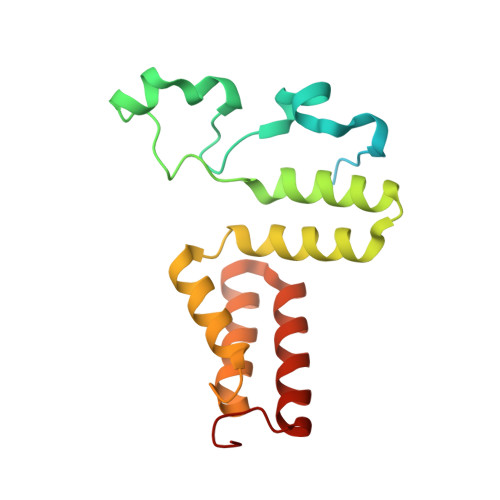
Redesign of coenzyme B(12) dependent diol dehydratase to be resistant to the mechanism-based inactivation by glycerol and act on longer chain 1,2-diols
Yamanishi, M., Kinoshita, K., Fukuoka, M., Saito, T., Tanokuchi, A., Ikeda, Y., Obayashi, H., Mori, K., Shibata, N., Tobimatsu, T., Toraya, T.(2012) FEBS J 279: 793-804
- PubMed: 22221669
- DOI: https://doi.org/10.1111/j.1742-4658.2012.08470.x
- Primary Citation of Related Structures:
3AUJ - PubMed Abstract:
Coenzyme B(12) dependent diol dehydratase undergoes mechanism-based inactivation by glycerol, accompanying the irreversible cleavage of the coenzyme Co-C bond. Bachovchin et al. [Biochemistry16, 1082-1092 (1977)] reported that glycerol bound in the G(S) conformation, in which the pro-S-CH(2) OH group is oriented to the hydrogen-abstracting site, primarily contributes to the inactivation reaction. To understand the mechanism of inactivation by glycerol, we analyzed the X-ray structure of diol dehydratase complexed with cyanocobalamin and glycerol. Glycerol is bound to the active site preferentially in the same conformation as that of (S)-1,2-propanediol, i.e. in the G(S) conformation, with its 3-OH group hydrogen bonded to Serα301, but not to nearby Glnα336. k(inact) of the Sα301A, Qα336A and Sα301A/Qα336A mutants with glycerol was much smaller than that of the wild-type enzyme. k(cat) /k(inact) showed that the Sα301A and Qα336A mutants are substantially more resistant to glycerol inactivation than the wild-type enzyme, suggesting that Serα301 and Glnα336 are directly or indirectly involved in the inactivation. The degree of preference for (S)-1,2-propanediol decreased on these mutations. The substrate activities towards longer chain 1,2-diols significantly increased on the Sα301A/Qα336A double mutation, probably because these amino acid substitutions yield more space for accommodating a longer alkyl group on C3 of 1,2-diols. Database Structural data are available in the Protein Data Bank under the accession number 3AUJ. Structured digital abstract • Diol dehydrase gamma subunit, Diol dehydrase beta subunit and Diol dehydrase alpha subunit physically interact by X-ray crystallography (View interaction).
Organizational Affiliation:
Department of Bioscience and Biotechnology, Graduate School of Natural Science and Technology, Okayama University, Japan.