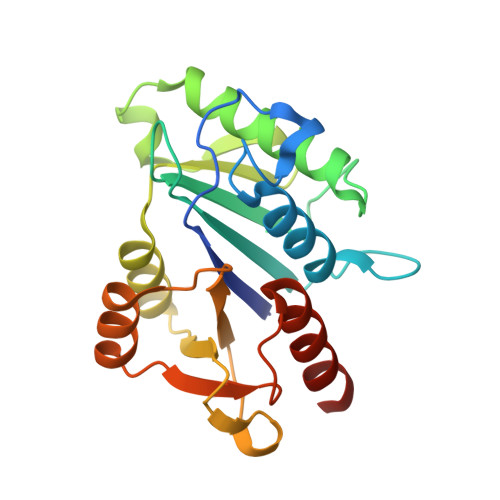
Crystal structure of Mj1640/DUF358 protein reveals a putative SPOUT-class RNA methyltransferase
Chen, H.Y., Yuan, Y.A.(2010) J Mol Cell Biol 2: 366-374
- PubMed: 21098051
- DOI: https://doi.org/10.1093/jmcb/mjq034
- Primary Citation of Related Structures:
3AI9, 3AIA - PubMed Abstract:
The proteins in DUF358 family are all bacterial proteins, which are ∼200 amino acids long with unknown function. Bioinformatics analysis suggests that these proteins contain several conserved arginines and aspartates that might adopt SPOUT-class fold. Here we report crystal structure of Methanocaldococcus jannaschii DUF358/Mj1640 in complex with S-adenosyl-L-methionine (SAM) at 1.4 Å resolution. The structure reveals a single domain structure consisting of eight-stranded β-sheets sandwiched by six α-helices at both sides. Similar to other SPOUT-class members, Mj1640 contains a typical deep trefoil knot at its C-terminus to accommodate the SAM cofactor. However, Mj1640 has limited structural extension at its N-terminus, which is unique to this family member. Mj1640 forms a dimer, which is mediated by two parallel pairs of α-helices oriented almost perpendicular to each other. Although Mj1640 shares close structural similarity with Nep1, the significant differences in N-terminal extension domain and the overall surface charge distribution strongly suggest that Mj1640 might target a different RNA sequence. Detailed structural analysis of the SAM-binding pocket reveals that Asp157 or Glu183 from its own monomer or Ser43 from the associate monomer probably plays the catalytic role for RNA methylation.
Organizational Affiliation:
Mechanobiology Institute, National University of Singapore, T-lab Building, 5A Engineering Drive 1, Singapore 117411, Singapore.