Structural determinant for switching between the polymerase and exonuclease modes in the PCNA-replicative DNA polymerase complex
Nishida, H., Mayanagi, K., Kiyonari, S., Sato, Y., Ishino, Y., Morikawa, K.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase | 775 | Pyrococcus furiosus | Mutation(s): 0 Gene Names: pol, PF0212 EC: 2.7.7.7 | 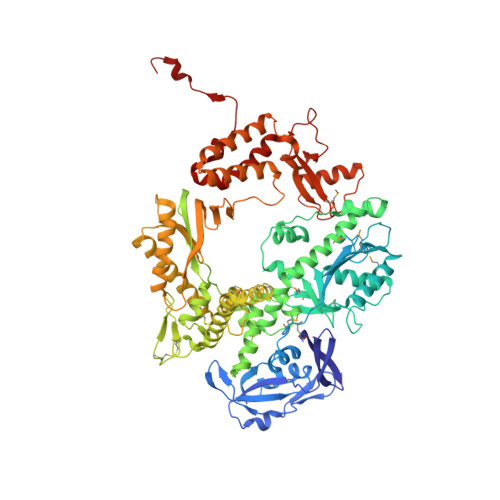 | |
UniProt | |||||
Find proteins for P61875 (Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)) Explore P61875 Go to UniProtKB: P61875 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P61875 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| DNA polymerase sliding clamp | 248 | Pyrococcus furiosus | Mutation(s): 3 Gene Names: pcn, PF0983 | 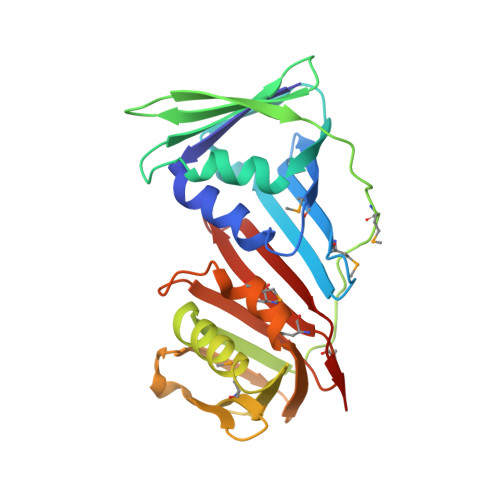 | |
UniProt | |||||
Find proteins for O73947 (Pyrococcus furiosus (strain ATCC 43587 / DSM 3638 / JCM 8422 / Vc1)) Explore O73947 Go to UniProtKB: O73947 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | O73947 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 77.354 | α = 90 |
| b = 90.451 | β = 90 |
| c = 186.211 | γ = 90 |
| Software Name | Purpose |
|---|---|
| CNS | refinement |
| HKL-2000 | data collection |
| HKL-2000 | data reduction |
| SCALEPACK | data scaling |
| SOLVE | phasing |