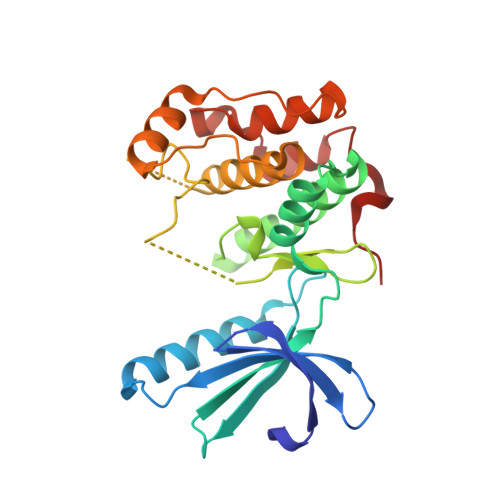
Aminopyrazine Inhibitors Binding to an Unusual Inactive Conformation of the Mitotic Kinase Nek2: Sar and Structural Characterization.
Whelligan, D.K., Solanki, S., Taylor, D., Thomson, D.W., Cheung, K.M., Boxall, K., Mas-Droux, C., Barillari, C., Burns, S., Grummitt, C.G., Collins, I., Van Montfort, R.L., Aherne, G.W., Bayliss, R., Hoelder, S.(2010) J Med Chem 53: 7682
- PubMed: 20936789
- DOI: https://doi.org/10.1021/jm1008727
- Primary Citation of Related Structures:
2XK3, 2XK4, 2XK6, 2XK7, 2XK8, 2XKC, 2XKD, 2XKE, 2XKF - PubMed Abstract:
We report herein the first systematic exploration of inhibitors of the mitotic kinase Nek2. Starting from HTS hit aminopyrazine 2, compounds with improved activity were identified using structure-based design. Our structural biology investigations reveal two notable observations. First, 2 and related compounds bind to an unusual, inactive conformation of the kinase which to the best of our knowledge has not been reported for other types of kinase inhibitors. Second, a phenylalanine residue at the center of the ATP pocket strongly affects the ability of the inhibitor to bind to the protein. The implications of these observations are discussed, and the work described here defines key features for potent and selective Nek2 inhibition, which will aid the identification of more advanced inhibitors of Nek2.
Organizational Affiliation:
Cancer Research UK Cancer Therapeutics Unit, The Institute of Cancer Research, 15 Cotswold Road, Sutton, Surrey SM2 5NG, United Kingdom.