Structure of Signal-Regulatory Protein Alpha: A Link to Antigen Receptor Evolution.
Hatherley, D., Graham, S.C., Harlos, K., Stuart, D.I., Barclay, A.N.(2009) J Biol Chem 284: 26613
- PubMed: 19628875
- DOI: https://doi.org/10.1074/jbc.M109.017566
- Primary Citation of Related Structures:
2WNG - PubMed Abstract:
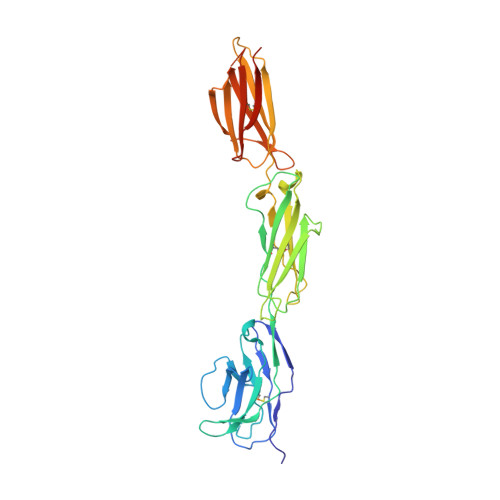
Signal-regulatory protein alpha (SIRPalpha) is a myeloid membrane receptor that interacts with the membrane protein CD47, a marker of self. We have solved the structure of the complete extracellular portion of SIRPalpha, comprising three immunoglobulin superfamily domains, by x-ray crystallography to 2.5 A resolution. These data, together with previous data on the N-terminal domain and its ligand CD47 (possessing a single immunoglobulin superfamily domain), show that the CD47-SIRPalpha interaction will span a distance of around 14 nm between interacting cells, comparable with that of an immunological synapse. The N-terminal (V-set) domain mediates binding to CD47, and the two others are found to be constant (C1-set) domains. C1-set domains are restricted to proteins involved in vertebrate antigen recognition: T cell antigen receptors, immunoglobulins, major histocompatibility complex antigens, tapasin, and beta2-microglobulin. The domains of SIRPalpha (domains 2 and 3) are structurally more similar to C1-set domains than any cell surface protein not involved in antigen recognition. This strengthens the suggestion from sequence analysis that SIRP is evolutionarily closely related to antigen recognition proteins.
Organizational Affiliation:
Sir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, United Kingdom.