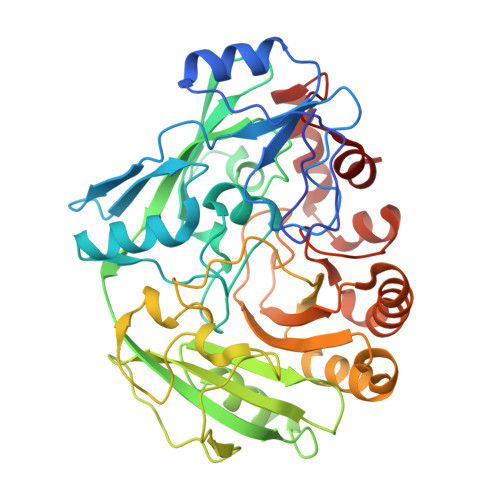
Structural Analysis of the Catalytic Mechanism and Stereoselectivity in Streptomyces Coelicolor Alditol Oxidase.
Forneris, F., Heuts, D.P.H.M., Delvecchio, M., Rovida, S., Fraaije, M.W., Mattevi, A.(2008) Biochemistry 47: 978
- PubMed: 18154360
- DOI: https://doi.org/10.1021/bi701886t
- Primary Citation of Related Structures:
2VFR, 2VFS, 2VFT, 2VFU, 2VFV - PubMed Abstract:
Alditol oxidase (AldO) from Streptomyces coelicolor A3(2) is a soluble monomeric flavin-dependent oxidase that performs selective oxidation of the terminal primary hydroxyl group of several alditols. Here, we report the crystal structure of the recombinant enzyme in its native state and in complex with both six-carbon (mannitol and sorbitol) and five-carbon substrates (xylitol). AldO shares the same folding topology of the members of the vanillyl-alcohol oxidase family of flavoenzymes and exhibits a covalently linked FAD which is located at the bottom of a funnel-shaped pocket that forms the active site. The high resolution of the three-dimensional structures highlights a well-defined hydrogen-bonding network that tightly constrains the substrate in the productive conformation for catalysis. Substrate binding occurs through a lock-and-key mechanism and does not induce conformational changes with respect to the ligand-free protein. A network of charged residues is proposed to favor catalysis through stabilization of the deprotonated form of the substrate. A His side chain acts as back door that "pushes" the substrate-reactive carbon atom toward the N5-C4a locus of the flavin. Analysis of the three-dimensional structure reveals possible pathways for diffusion of molecular oxygen and a small cavity on the re side of the flavin that may host oxygen during FAD reoxidation. These features combined with the tight shape of the catalytic site provide insights into the mechanism of AldO-mediated regioselective oxidation reactions and its substrate specificity.
Organizational Affiliation:
Department of Genetics and Microbiology, University of Pavia, Via Ferrata 1, 27100 Pavia, Italy.