Selected Mutations in Bacillus Subtilis Levansucrase Semi-Conserved Regions Affecting its Biochemical Properties
Ortiz-Soto, M.E., Rivera, M., Rudino-Pinera, E., Olvera, C., Lopez-Munguia, A.(2008) Protein Eng Des Sel 21: 589-595
- PubMed: 18596022
- DOI: https://doi.org/10.1093/protein/gzn036
- Primary Citation of Related Structures:
2VDT - PubMed Abstract:
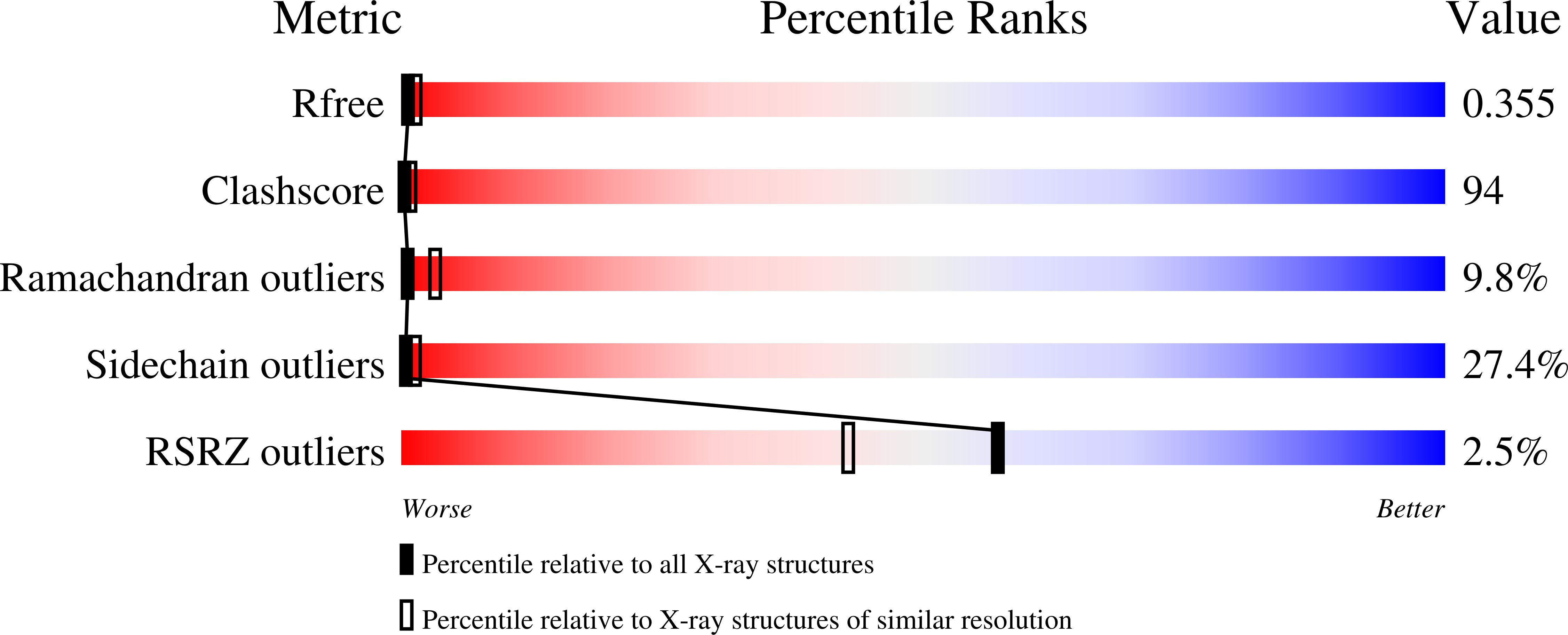
Levansucrases (LS) are fructosyltransferases (FTFs) belonging to family 68 of glycoside hydrolases (GH68) using sucrose as substrate to synthesize levan, a fructose polymer. From a multiple sequence analysis of GH68 family proteins, nine residues were selected and their role in acceptor and product specificity, as well as in biochemical Bacillus subtilis LS properties, was investigated. A product specificity modification was obtained with mutants Y429N and R433A that no longer produce levan but exclusively oligosaccharides. An effect of the mutation S164A was observed on enzyme stability and kinetic behavior; this mutation also induces a levan activation effect that enhances the reaction rate. We report the crystallographic structure of this mutant and found that S164 is an important residue to maintain the nucleophile position in the active site. We also found evidence of the important role of Y429 in acceptor specificity: this is a key residue coordinating the sucrose position in the catalytic domain-binding pocket. Some of these mutations resulted in LS with a broad range of specificities and new biochemical properties.
Organizational Affiliation:
Departamento de Ingeniería Celular y Biocatálisis, Instituto de Biotecnología, UNAM, Cuernavaca, Morelos 62210, Mexico.