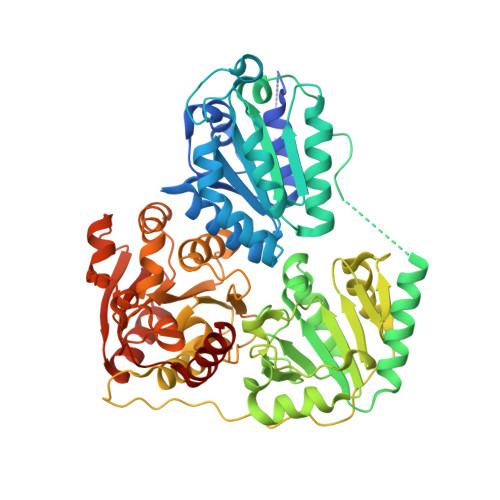
Crystal Structure of the Branched-Chain Keto Acid Decarboxylase (Kdca) from Lactococcus Lactis Provides Insights Into the Structural Basis for the Chemo- and Enantioselective Carboligation Reaction
Berthold, C.L., Gocke, D., Wood, M.D., Leeper, F., Pohl, M., Schneider, G.(2007) Acta Crystallogr D Biol Crystallogr 63: 1217
- PubMed: 18084069
- DOI: https://doi.org/10.1107/S0907444907050433
- Primary Citation of Related Structures:
2VBF, 2VBG - PubMed Abstract:
The thiamin diphosphate (ThDP) dependent branched-chain keto acid decarboxylase (KdcA) from Lactococcus lactis catalyzes the decarboxylation of 3-methyl-2-oxobutanoic acid to 3-methylpropanal (isobutyraldehyde) and CO2. The enzyme is also able to catalyze carboligation reactions with an exceptionally broad substrate range, a feature that makes KdcA a potentially valuable biocatalyst for C-C bond formation, in particular for the enzymatic synthesis of diversely substituted 2-hydroxyketones with high enantioselectivity. The crystal structures of recombinant holo-KdcA and of a complex with an inhibitory ThDP analogue mimicking a reaction intermediate have been determined to resolutions of 1.6 and 1.8 A, respectively. KdcA shows the fold and cofactor-protein interactions typical of thiamin-dependent enzymes. In contrast to the tetrameric assembly displayed by most other ThDP-dependent decarboxylases of known structure, KdcA is a homodimer. The crystal structures provide insights into the structural basis of substrate selectivity and stereoselectivity of the enzyme and thus are suitable as a framework for the redesign of the substrate profile in carboligation reactions.
Organizational Affiliation:
Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-17177 Stockholm, Sweden.