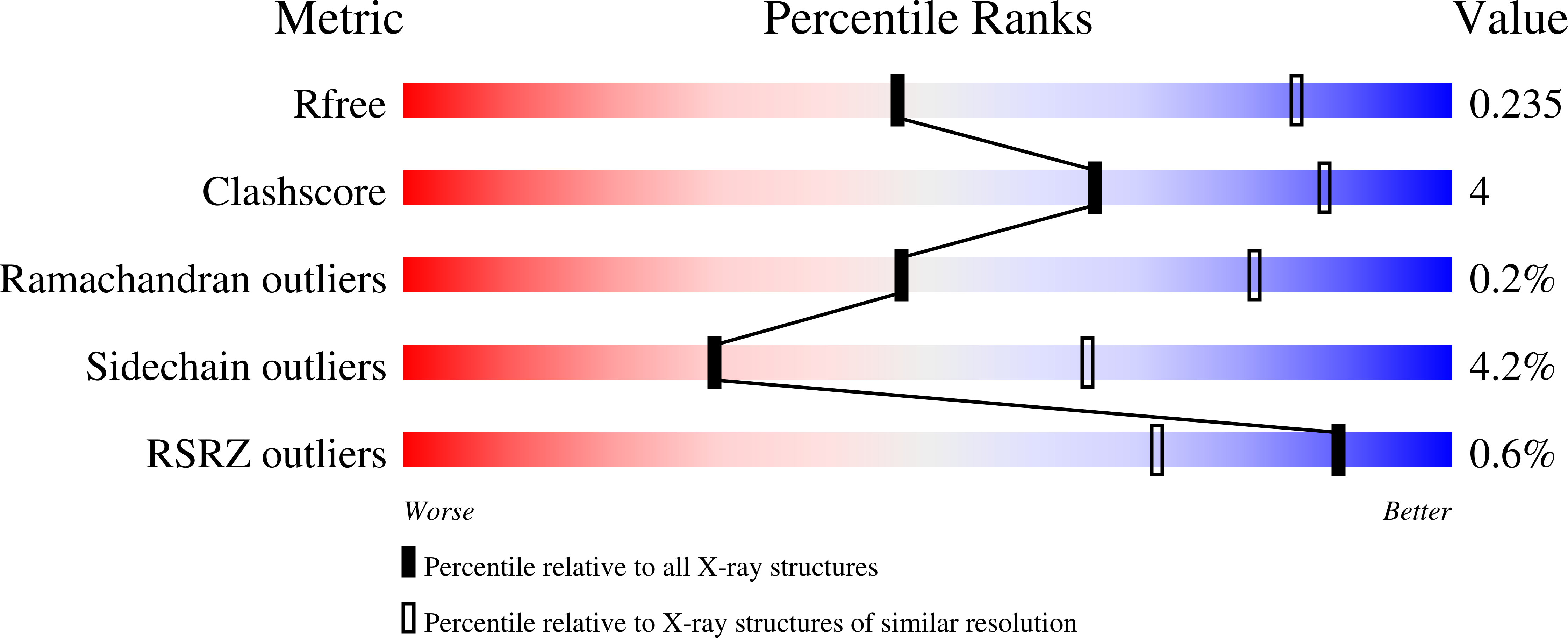
Crystal structures of Arabidopsis thaliana cell-wall invertase mutants in complex with sucrose.
Lammens, W., Le Roy, K., Van Laere, A., Rabijns, A., Van den Ende, W.(2008) J Mol Biol 377: 378-385
- PubMed: 18258263
- DOI: https://doi.org/10.1016/j.jmb.2007.12.074
- Primary Citation of Related Structures:
2QQU, 2QQV, 2QQW - PubMed Abstract:
In plants, cell-wall invertases fulfil important roles in carbohydrate partitioning, growth, development and crop yield. In this study, we report on different X-ray crystal structures of Arabidopsis thaliana cell-wall invertase 1 (AtcwINV1) mutants with sucrose. These structures reveal a detailed view of sucrose binding in the active site of the wild-type AtcwINV1. Compared to related enzyme-sucrose complexes, important differences in the orientation of the glucose subunit could be observed. The structure of the E203Q AtcwINV1 mutant showed a complete new binding modus, whereas the D23A, E203A and D239A structures most likely represent the productive binding modus. Together with a hydrophobic zone formed by the conserved W20, W47 and W82, the residues N22, D23, R148, E203, D149 and D239 are necessary to create the ideal sucrose-binding pocket. D239 can interact directly with the glucose moiety of sucrose, whereas K242 has an indirect role in substrate stabilization. Most probably, K242 keeps D239 in a favourable position upon substrate binding. Unravelling the exact position of sucrose in plant cell-wall invertases is a necessary step towards the rational design of superior invertases to further increase crop yield and biomass production.
Organizational Affiliation:
Laboratorium voor Moleculaire Plantenfysiologie, Faculteit Wetenschappen, Departement Biologie, K.U. Leuven, Kasteelpark Arenberg 31, bus 2434, B-3001 Heverlee, Belgium.