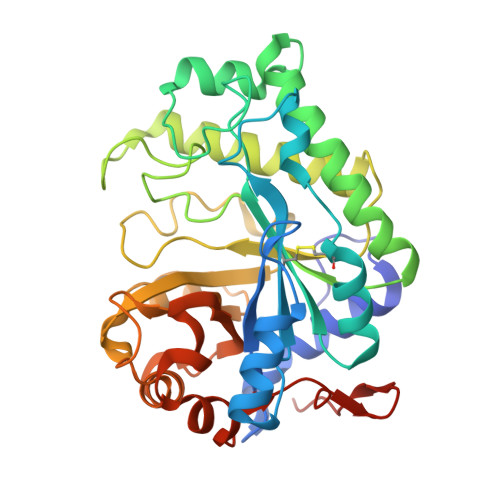
From Structure to Function: Insights into the Catalytic Substrate Specificity and Thermostability Displayed by Bacillus subtilis Mannanase BCman
Yan, X.X., An, X.M., Gui, L.L., Liang, D.C.(2008) J Mol Biol 379: 535-544
- PubMed: 18455734
- DOI: https://doi.org/10.1016/j.jmb.2008.03.068
- Primary Citation of Related Structures:
2QHA - PubMed Abstract:
BCman, a beta-mannanase from the plant root beneficial bacterium Bacillus subtilis Z-2, has a potential to be used in the production of mannooligosaccharide, which shows defense induction activity on both melon and tobacco, and plays an important role in the biological control of plant disease. Here we report the biochemical properties and crystal structure of BCman-GH26 enzyme. Kinetic analysis reveals that BCman is an endo-beta-mannanase, specific for mannan, and has no activity on mannooligosaccharides. The catalytic acid/base Glu167 and nucleophile Glu266 are positioned on the beta4 and beta7 strands, respectively. The 1.45-A crystal structure reveals that BCman is a typical (beta/alpha)(8) folding type. One large difference from the saddle-shaped active center of other endo-beta-mannanases is the presence of a shallow-dish-shaped active center and substrate-binding site that are both unique to BCman. These differences are mainly due to important changes in the length and position of loop 1 (Phe37-Met47), loop 2 (Ser103-Ala134), loop3 (Phe162-Asn185), loop 4 (Tyr215-Ile236), loop 5 (Pro269-Tyr278), and loop 6 (Trp298-Gly309), all of which surround the active site. Data from isothermal titration calorimetry and crystallography indicated only two substrate-binding subsites (+1 and -1) within the active site of BCman. These two sites are involved in the enzyme's mannan degradation activity and in restricting the binding capacity for mannooligosaccharides. Binding and catalysis of BCman to mannan is mediated mainly by a surface containing a strip of solvent-exposed aromatic rings of Trp302, Trp298, Trp172, and Trp72. Additionally, BCman contains a disulfide bond (Cys66Cys86) and a special His1-His23-Glu336 metal-binding site. This secondary structure is a key factor in the enzyme's stability.
Organizational Affiliation:
National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, P. R. China.