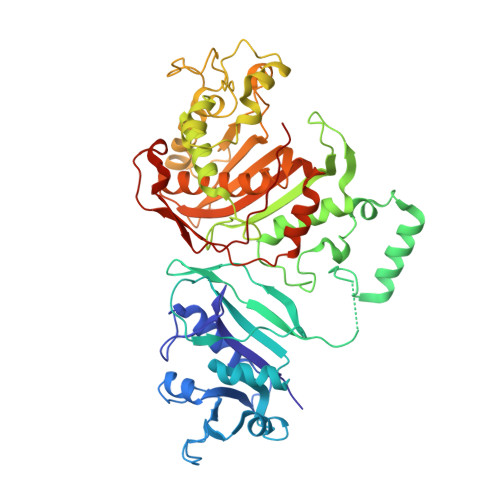
Nonconserved residues Ala287 and Ser290 of the Cryptosporidium hominis thymidylate synthase domain facilitate its rapid rate of catalysis
Doan, L.T., Martucci, W.E., Vargo, M.A., Atreya, C.E., Anderson, K.S.(2007) Biochemistry 46: 8379-8391
- PubMed: 17580969
- DOI: https://doi.org/10.1021/bi700531r
- Primary Citation of Related Structures:
2OIP - PubMed Abstract:
Cryptosporidium hominis TS-DHFR exhibits an unusually high rate of catalysis at the TS domain, at least 10-fold greater than those of other TS enzymes. Using site-directed mutagenesis, we have mutated residues Ala287 and Ser290 in the folate-binding helix to phenylalanine and glycine, respectively, the corresponding residues in human and most other TS enzymes. Our results show that the mutant A287F, the mutant S290G, and the double mutant all have reduced affinities for methylene tetrahydrofolate and reduced rates of reaction at the TS domain. Interestingly, the S290G mutant enzyme had the lowest TS activity, with a catalytic efficiency approximately 200-fold lower than that of the wild type (WT). The rate of conformational change of the S290G mutant is approximately 80 times slower than that of WT, resulting in a change in the rate-limiting step from hydride transfer to covalent ternary complex formation. We have determined the crystal structure of ligand-bound S290G mutant enzyme, which shows that the primary effect of the mutation is an increase in the distance between the TS ligands. The kinetic and crystal structure data presented here provide the first evidence explaining the unusually fast TS rate in C. hominis.
Organizational Affiliation:
Department of Pharmacology, Yale University School of Medicine, 333 Cedar Street, New Haven, Connecticut 06520, USA.