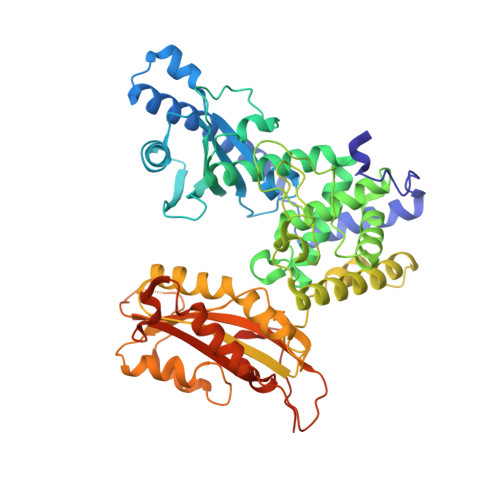
X-ray crystallographic and steady state fluorescence characterization of the protein dynamics of yeast polyadenylate polymerase.
Balbo, P.B., Toth, J., Bohm, A.(2007) J Mol Biol 366: 1401-1415
- PubMed: 17223131
- DOI: https://doi.org/10.1016/j.jmb.2006.12.030
- Primary Citation of Related Structures:
2HHP, 2O1P - PubMed Abstract:
Polyadenylate polymerase (PAP) catalyzes the synthesis of poly(A) tails on the 3'-end of pre-mRNA. PAP is composed of three domains: an N-terminal nucleotide-binding domain (homologous to the palm domain of DNA and RNA polymerases), a middle domain (containing other conserved, catalytically important residues), and a unique C-terminal domain (involved in protein-protein interactions required for 3'-end formation). Previous X-ray crystallographic studies have shown that the domains are arranged in a V-shape such that they form a central cleft with the active site located at the base of the cleft at the interface between the N-terminal and middle domains. In the previous studies, the nucleotides were bound directly to the N-terminal domain and exhibited a conspicuous lack of adenine-specific interactions that would constitute nucleotide recognition. Furthermore, it was postulated that base-specific contacts with residues in the middle domain could occur either as a result of a change in the conformation of the nucleotide or domain movement. To address these issues and to better characterize the structural basis of substrate recognition and catalysis, we report two new crystal structures of yeast PAP. A comparison of these structures reveals that the N-terminal and C-terminal domains of PAP move independently as rigid bodies along two well defined axes of rotation. Modeling of the nucleotide into the most closed state allows us to deduce specific nucleotide interactions involving residues in the middle domain (K215, Y224 and N226) that are proposed to be involved in substrate binding and specificity. To further investigate the nature of PAP domain flexibility, 2-aminopurine labeled molecular probes were employed in steady state fluorescence and acrylamide quenching experiments. The results suggest that the closed domain conformation is stabilized upon recognition of the correct subtrate, MgATP, in an enzyme-substrate ternary complex. The implications of these results on the enzyme mechanism of PAP and the possible role for domain motion in an induced fit mechanism are discussed.
Organizational Affiliation:
Tufts University School of Medicine, Sackler School of Graduate Biomedical Sciences, Department of Biochemistry, Boston, MA 02111, USA.