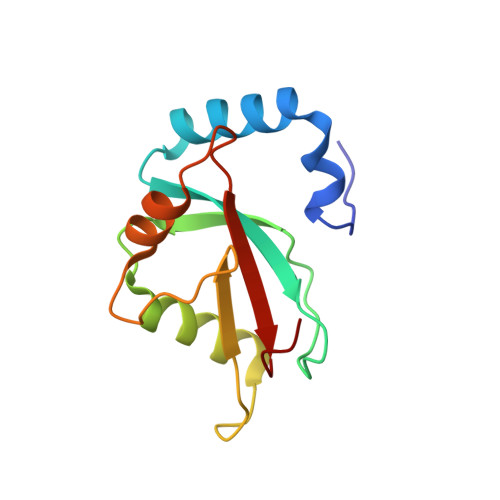
Characterization of the Interaction of GABARAPL-1 with the LIR Motif of NBR1.
Rozenknop, A., Rogov, V.V., Rogova, N.Y., Lohr, F., Guntert, P., Dikic, I., Dotsch, V.(2011) J Mol Biol 410: 477-487
- PubMed: 21620860
- DOI: https://doi.org/10.1016/j.jmb.2011.05.003
- Primary Citation of Related Structures:
2L8J - PubMed Abstract:
Selective autophagy requires the specific segregation of targeted proteins into autophagosomes. The selectivity is mediated by autophagy receptors, such as p62 and NBR1, which can bind to autophagic effector proteins (Atg8 in yeast, MAP1LC3 protein family in mammals) anchored in the membrane of autophagosomes. Recognition of autophagy receptors by autophagy effectors takes place through an LC3 interaction region (LIR). The canonical LIR motif consists of a WXXL sequence, N-terminally preceded by negatively charged residues. The LIR motif of NBR1 presents differences to this classical LIR motif with a tyrosine residue and an isoleucine residue substituting the tryptophan residue and the leucine residue, respectively. We have determined the structure of the GABARAPL-1/NBR1-LIR complex and studied the influence of the different residues belonging to the LIR motif for the interaction with several mammalian autophagy modifiers (LC3B and GABARAPL-1). Our results indicate that the presence of a tryptophan residue in the LIR motif increases the binding affinity. Substitution by other aromatic amino acids or increasing the number of negatively charged residues at the N-terminus of the LIR motif, however, has little effect on the binding affinity due to enthalpy-entropy compensation. This indicates that different LIRs can interact with autophagy modifiers with unique binding properties.
Organizational Affiliation:
Institute of Biophysical Chemistry and Center for Biomolecular Magnetic Resonance, Goethe University, 60438 Frankfurt, Germany.