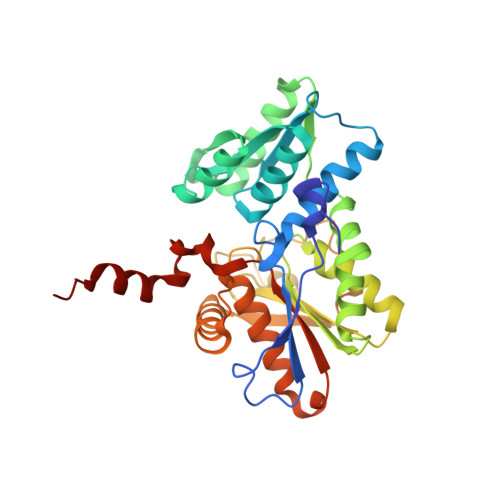
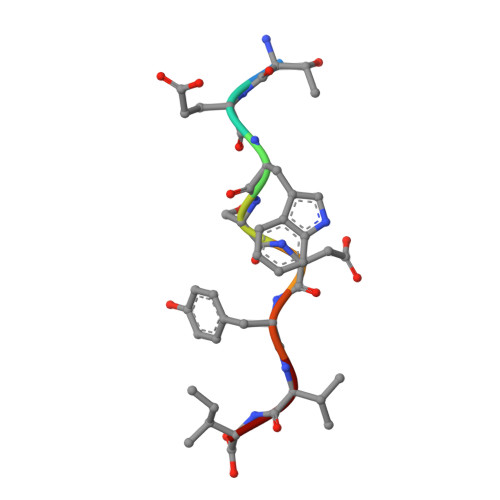
Structural basis for interaction of o-acetylserine sulfhydrylase and serine acetyltransferase in the Arabidopsis cysteine synthase complex.
Francois, J.A., Kumaran, S., Jez, J.M.(2006) Plant Cell 18: 3647-3655
- PubMed: 17194764
- DOI: https://doi.org/10.1105/tpc.106.047316
- Primary Citation of Related Structures:
2ISQ - PubMed Abstract:
In plants, association of O-acetylserine sulfhydrylase (OASS) and Ser acetyltransferase (SAT) into the Cys synthase complex plays a regulatory role in sulfur assimilation and Cys biosynthesis. We determined the crystal structure of Arabidopsis thaliana OASS (At-OASS) bound with a peptide corresponding to the C-terminal 10 residues of Arabidopsis SAT (C10 peptide) at 2.9-A resolution. Hydrogen bonding interactions with key active site residues (Thr-74, Ser-75, and Gln-147) lock the C10 peptide in the binding site. C10 peptide binding blocks access to OASS catalytic residues, explaining how complex formation downregulates OASS activity. Comparison with bacterial OASS suggests that structural plasticity in the active site allows binding of SAT C termini with dissimilar sequences at structurally similar OASS active sites. Calorimetric analysis of the effect of active site mutations (T74S, S75A, S75T, and Q147A) demonstrates that these residues are important for C10 peptide binding and that changes at these positions disrupt communication between active sites in the homodimeric enzyme. We also demonstrate that the C-terminal Ile of the C10 peptide is required for molecular recognition by At-OASS. These results provide new insights into the molecular mechanism underlying formation of the Cys synthase complex and provide a structural basis for the biochemical regulation of Cys biosynthesis in plants.
Organizational Affiliation:
Donald Danforth Plant Science Center, St. Louis, Missouri 63132, USA.