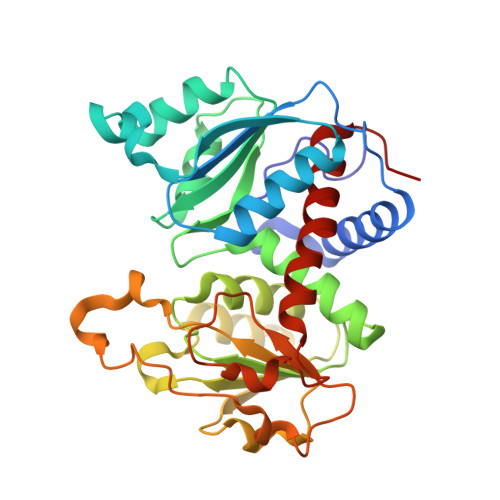
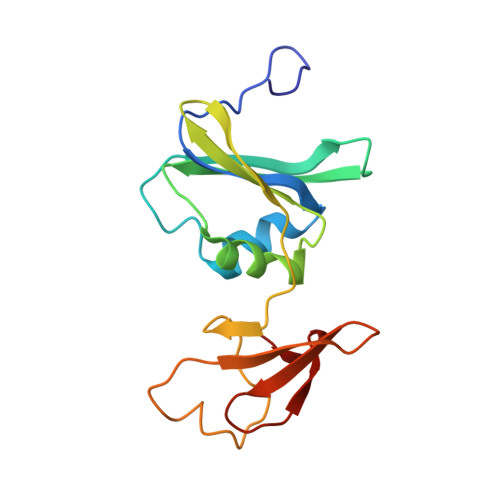
Use of L-asparagine and N-phosphonacetyl-L-asparagine to investigate the linkage of catalysis and homotropic cooperativity in E. coli aspartate transcarbomoylase.
Cardia, J.P., Eldo, J., Xia, J., O'Day, E.M., Tsuruta, H., Gryncel, K.R., Kantrowitz, E.R.(2008) Proteins 71: 1088-1096
- PubMed: 18004787
- DOI: https://doi.org/10.1002/prot.21760
- Primary Citation of Related Structures:
2IPO - PubMed Abstract:
The mechanism of domain closure and the allosteric transition of Escherichia coli aspartate transcarbamoylase (ATCase) are investigated using L-Asn, in the presence of carbamoyl phosphate (CP), and N-phosphonacetyl-L-asparagine (PASN). ATCase was found to catalyze the carbamoylation of L-Asn with a K(m) of 122 mM and a maximal velocity 10-fold lower than observed with the natural substrate, L-Asp. As opposed to L-Asp, no cooperativity was observed with respect to L-Asn. Time-resolved small-angle X-ray scattering (SAXS) and fluorescence experiments revealed that the combination of CP and L-Asn did not convert the enzyme from the T to the R state. PASN was found to be a potent inhibitor of ATCase exhibiting a K(D) of 8.8 microM. SAXS experiments showed that PASN was able to convert the entire population of molecules to the R state. Analysis of the crystal structure of the enzyme in the presence of PASN revealed that the binding of PASN was similar to that of the R-state complex of ATCase with N-phosphonaceyl-L-aspartate, another potent inhibitor of the enzyme. The linking of CP and L-Asn into one molecule, PASN, correctly orients the asparagine moiety in the active site to induce domain closure and the allosteric transition. This entropic effect allows for the high affinity binding of PASN. However, the binding of L-Asn, in the presence of a saturating concentration of CP, does not induce the closure of the two domains of the catalytic chain, nor does the enzyme undergo the transition to the high-activity high- affinity R structure. These results imply that Arg229, which interacts with the beta-carboxylate of L-Asp, plays a critical role in the orientation of L-Asp in the active site and demonstrates the requirement of the beta-carboxylate of L-Asp in the mechanism of domain closure and the allosteric transition in E. coli ATCase.
Organizational Affiliation:
Department of Chemistry, Merkert Chemistry Center, Boston College, Chestnut Hill, Massachusetts 02467-3807, USA.