Combinatorial optimization of a CD4-mimetic miniprotein and cocrystal structures with HIV-1 gp120 envelope glycoprotein.
Stricher, F., Huang, C.C., Descours, A., Duquesnoy, S., Combes, O., Decker, J.M., Kwon, Y.D., Lusso, P., Shaw, G.M., Vita, C., Kwong, P.D., Martin, L.(2008) J Mol Biol 382: 510-524
- PubMed: 18619974
- DOI: https://doi.org/10.1016/j.jmb.2008.06.069
- Primary Citation of Related Structures:
2I5Y, 2I60 - PubMed Abstract:
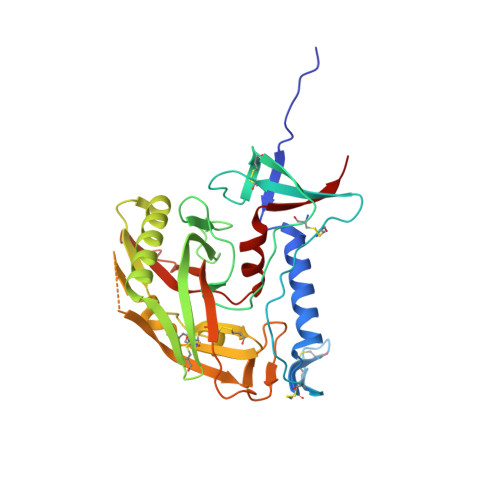
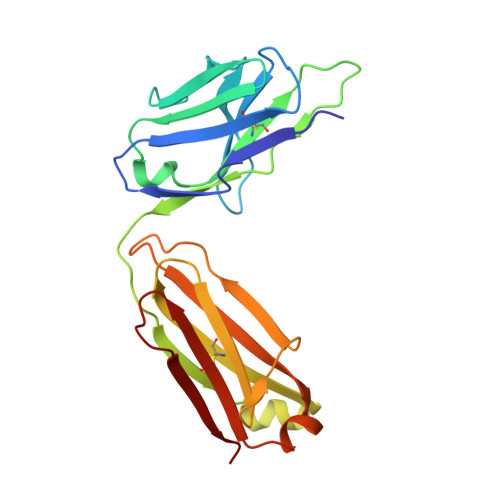
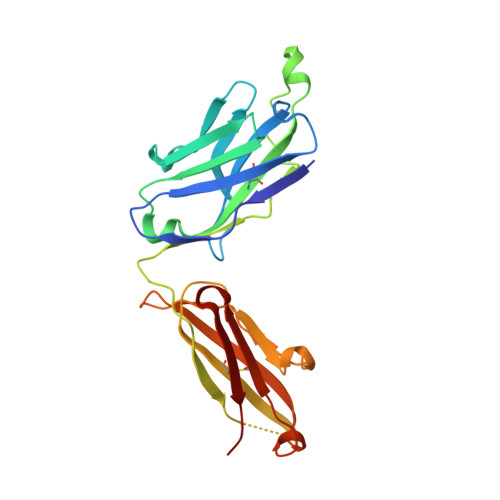
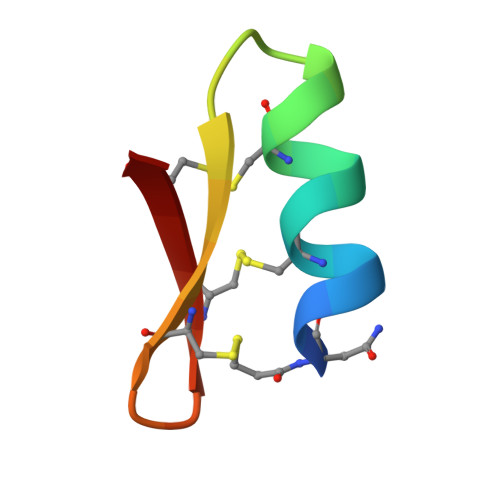
Miniproteins provide a bridge between proteins and small molecules. Here we adapt methods from combinatorial chemistry to optimize CD4M33, a synthetic miniprotein into which we had previously transplanted the HIV-1 gp120 binding surface of the CD4 receptor. Iterative deconvolution of generated libraries produced CD4M47, a derivative of CD4M33 that had been optimized at four positions. Surface plasmon resonance demonstrated fourfold to sixfold improvement in CD4M47 affinity for gp120 to a level about threefold tighter than that of CD4 itself. Assessment of the neutralization properties of CD4M47 against a diverse range of isolates spanning from HIV-1 to SIVcpz showed that CD4M47 retained the extraordinary breadth of the parent CD4M33, but yielded only limited improvements in neutralization potencies. Crystal structures of CD4M47 and a phenylalanine variant ([Phe23]M47) were determined at resolutions of 2.4 and 2.6 A, in ternary complexes with HIV-1 gp120 and the 17b antibody. Analysis of these structures revealed a correlation between mimetic affinity for gp120 and overall mimetic-gp120 interactive surface. A correlation was also observed between CD4- and mimetic-induced gp120 structural similarity and CD4- and mimetic-induced gp120 affinity for the CCR5 coreceptor. Despite mimetic substitutions, including a glycine-to-(d)-proline change, the gp120 conformation induced by CD4M47 was as close or closer to the conformation induced by CD4 as the one induced by the parent CD4M33. Our results demonstrate the ability of combinatorial chemistry to optimize a disulfide-containing miniprotein, and of structural biology to decipher the resultant interplay between binding affinity, neutralization breadth, molecular mimicry, and induced affinity for CCR5.
Organizational Affiliation:
CEA, iBiTecS, SIMOPRO, Gif-sur-Yvette F-91191, France.