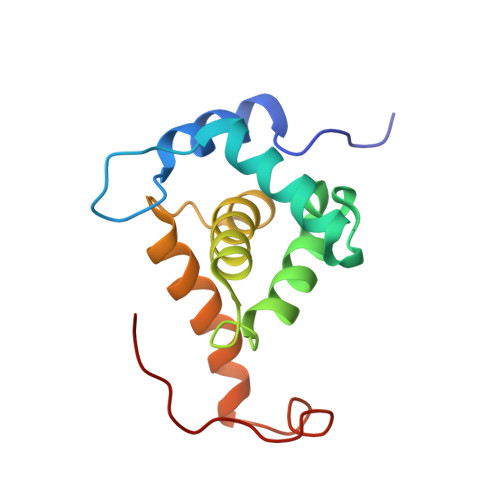
Structural basis for targeting HIV-1 Gag proteins to the plasma membrane for virus assembly.
Saad, J.S., Miller, J., Tai, J., Kim, A., Ghanam, R.H., Summers, M.F.(2006) Proc Natl Acad Sci U S A 103: 11364-11369
- PubMed: 16840558
- DOI: https://doi.org/10.1073/pnas.0602818103
- Primary Citation of Related Structures:
2H3F, 2H3I, 2H3Q, 2H3V, 2H3Z - PubMed Abstract:
During the late phase of HIV type 1 (HIV-1) replication, newly synthesized retroviral Gag proteins are targeted to the plasma membrane of most hematopoietic cell types, where they colocalize at lipid rafts and assemble into immature virions. Membrane binding is mediated by the matrix (MA) domain of Gag, a 132-residue polypeptide containing an N-terminal myristyl group that can adopt sequestered and exposed conformations. Although exposure is known to promote membrane binding, the mechanism by which Gag is targeted to specific membranes has yet to be established. Recent studies have shown that phosphatidylinositol (PI) 4,5-bisphosphate [PI(4,5)P(2)], a factor that regulates localization of cellular proteins to the plasma membrane, also regulates Gag localization and assembly. Here we show that PI(4,5)P(2) binds directly to HIV-1 MA, inducing a conformational change that triggers myristate exposure. Related phosphatidylinositides PI, PI(3)P, PI(4)P, PI(5)P, and PI(3,5)P(2) do not bind MA with significant affinity or trigger myristate exposure. Structural studies reveal that PI(4,5)P(2) adopts an "extended lipid" conformation, in which the inositol head group and 2'-fatty acid chain bind to a hydrophobic cleft, and the 1'-fatty acid and exposed myristyl group bracket a conserved basic surface patch previously implicated in membrane binding. Our findings indicate that PI(4,5)P(2) acts as both a trigger of the myristyl switch and a membrane anchor and suggest a potential mechanism for targeting Gag to membrane rafts.
Organizational Affiliation:
Howard Hughes Medical Institute, University of Maryland, Baltimore County, 1000 Hilltop Circle, Baltimore, MD 21250, USA.