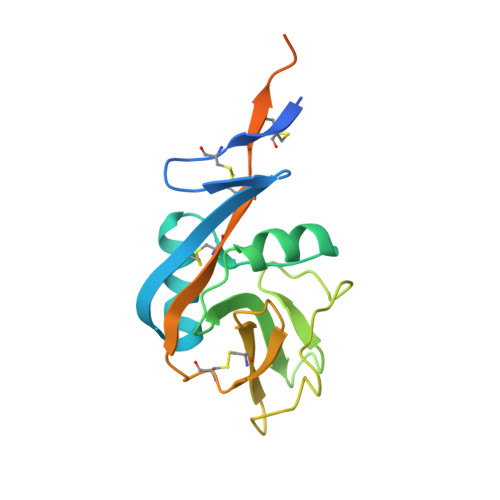
Structural Changes in the Lectin Domain of CD23, the Low-Affinity IgE Receptor, upon Calcium Binding.
Wurzburg, B.A., Tarchevskaya, S.S., Jardetzky, T.S.(2006) Structure 14: 1049-1058
- PubMed: 16765898
- DOI: https://doi.org/10.1016/j.str.2006.03.017
- Primary Citation of Related Structures:
2H2R, 2H2T - PubMed Abstract:
CD23, the low-affinity receptor for IgE (Fc epsilonRII), regulates IgE synthesis and also mediates IgE-dependent antigen transport and processing. CD23 is a unique Fc receptor belonging to the C-type lectin-like domain superfamily and binds IgE in an unusual, non-lectin-like manner, requiring calcium but not carbohydrate. We have solved the high-resolution crystal structures of the human CD23 lectin domain in the presence and absence of Ca2+. The crystal structures differ significantly from a previously determined NMR structure and show that calcium binding occurs at the principal binding site, but not at an auxiliary site that appears to be absent in human CD23. Conformational differences between the apo and Ca2+ bound structures suggest how IgE-Fc binding can be both calcium-dependent and carbohydrate-independent.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology and Cell Biology, Northwestern University, Evanston, Illinois 60208, USA.