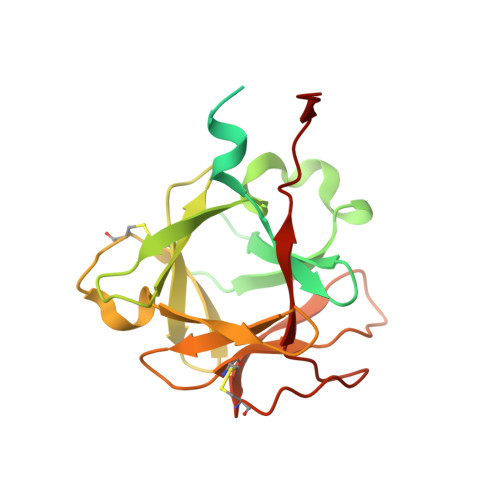
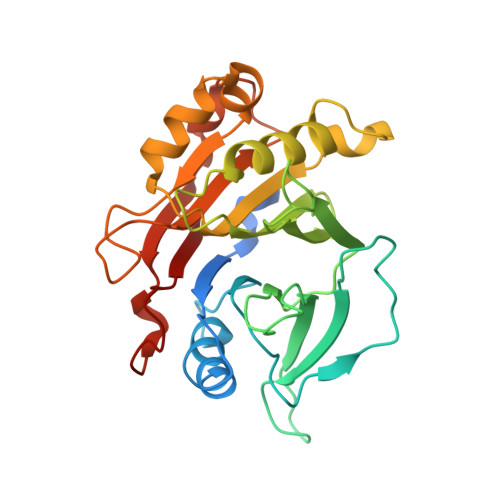
Variation of loop sequence alters stability of cytolethal distending toxin (CDT): crystal structure of CDT from Actinobacillus actinomycetemcomitans
Yamada, T., Komoto, J., Saiki, K., Konishi, K., Takusagawa, F.(2006) Protein Sci 15: 362-372
- PubMed: 16434747
- DOI: https://doi.org/10.1110/ps.051790506
- Primary Citation of Related Structures:
2F2F - PubMed Abstract:
Cytolethal distending toxin (CDT) secreted by Actinobacillus actinomycetemcomitans induces cell cycle arrest of cultured cells in the G2 phase. The crystal structure of the natural form of A. actinomycetemcomitans DCT (aCDT) has been determined at 2.4 A resolution. aCDT is a heterotrimer consisting of the three subunits, aCdtA, aCdtB, and aCdtC. Two crystallographically independent aCDTs form a dimer through interactions of the aCdtB subunits. The primary structure of aCDT has 94.3% identity with that of Haemophilus ducreyi CDT (hCDT), and the structure of aCDT is quite similar to that of hCDT reconstituted from the three subunits determined recently. However, the molecular packings in the crystal structures of aCDT and hCDT are quite different. A careful analysis of molecular packing suggests that variation of the amino acid residues in a nonconserved loop (181TSSPSSPERRGY192 of aCdtB and 181NSSSSPPERRVY192 of hCdtB) is responsible for the different oligomerization of very similar CDTs. The loop of aCdtB has a conformation to form a dimer, while the loop conformation of hCdtB prevents hCDT from forming a dimer. Although dimerization of aCDT does not affect toxic activity, it changes the stability of protein. aCDT rapidly aggregates and loses toxic activity in the absence of sucrose in a buffered solution, while hCDT is stable and retains toxic activity. Another analysis of crystal structures of aCDT and hCDT suggests that the receptor contact area is in the deep groove between CdtA and CdtC, and the characteristic "aromatic patch" on CdtA.
Organizational Affiliation:
Department of Molecular Biosciences, University of Kansas, Lawrence, KS 66045-7534, USA.