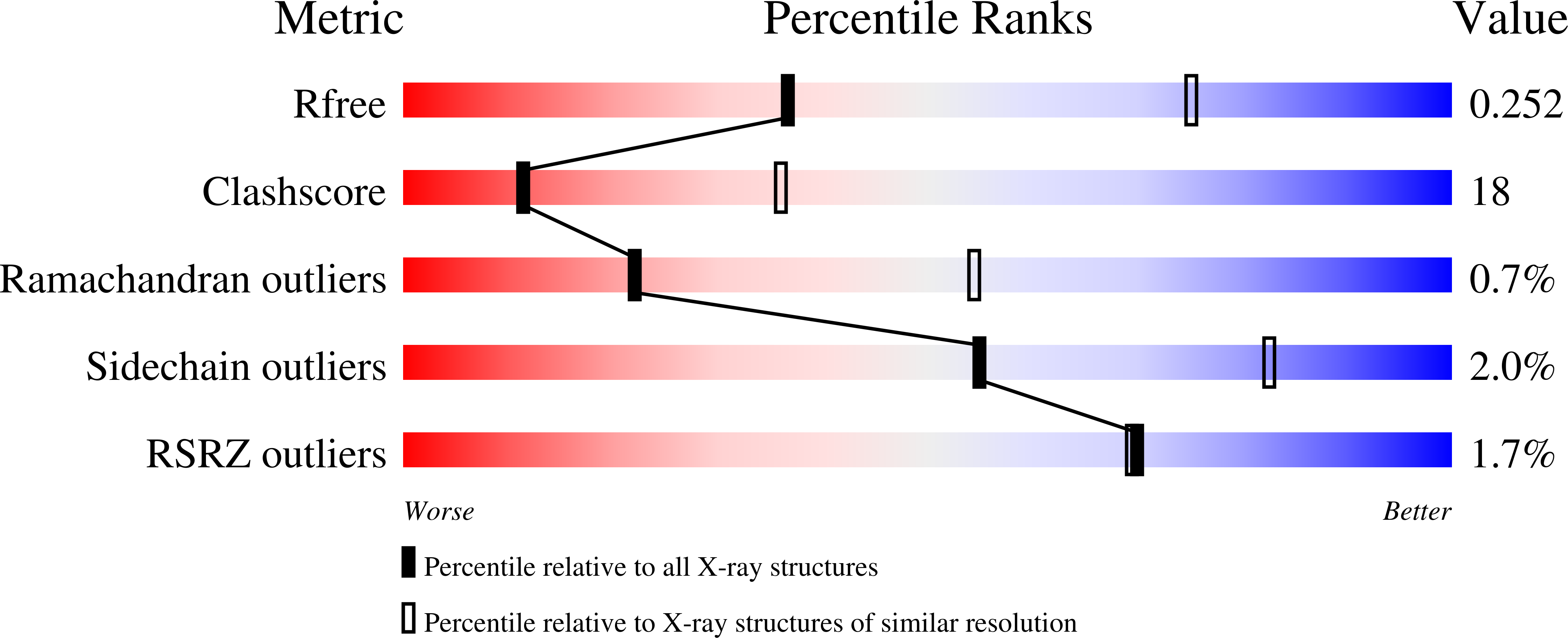
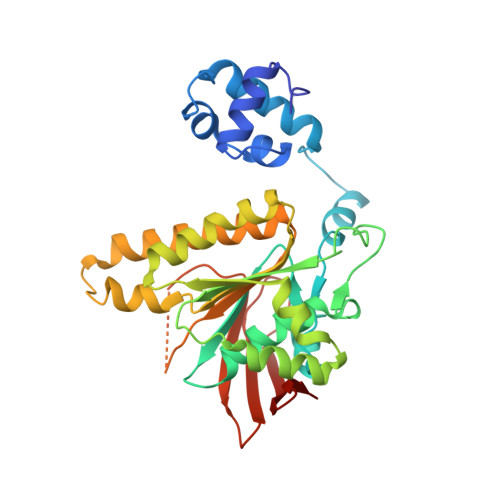
Asp302 determines potassium dependence of a RadA recombinase from Methanococcus voltae.
Qian, X., He, Y., Wu, Y., Luo, Y.(2006) J Mol Biol 360: 537-547
- PubMed: 16782126
- DOI: https://doi.org/10.1016/j.jmb.2006.05.058
- Primary Citation of Related Structures:
2F1H, 2F1I, 2F1J - PubMed Abstract:
Archaeal RadA/Rad51 are close homologues of eukaryal Rad51/DMC1. Such recombinases, as well as their bacterial RecA orthologues, form helical nucleoprotein filaments in which a hallmark strand exchange reaction occurs between homologous DNA substrates. Our recent ATPase and structure studies on RadA recombinase from Methanococcus voltae have suggested that not only magnesium but also potassium ions are absorbed at the ATPase center. Potassium, but not sodium, stimulates the ATP hydrolysis reaction with an apparent dissociation constant of approximately 40 mM. The minimal inhibitory effect by 40 mM NaCl further suggests that the protein does not have adequate affinity for sodium. The wild-type protein's strand exchange activity is also stimulated by potassium with an apparent dissociation constant of approximately 35 mM. We made site-directed mutations at the potassium-contacting residues Glu151 and Asp302. The mutant proteins are expectedly defective in promoting ATP hydrolysis. Similar potassium preference in strand exchange is observed for the E151D and E151K proteins. The D302K protein, however, shows comparable strand exchange efficiencies in the presence of either potassium or sodium. Crystallized E151D filaments reveal a potassium-dependent conformational change similar to what has previously been observed with the wild-type protein. We interpret these data as suggesting that both ATP hydrolysis and DNA strand exchange requires accessibility to an "active" conformation similar to the crystallized ATPase-active form in the presence of ATP, Mg2+ and K+.
Organizational Affiliation:
Department of Biochemistry, University of Saskatchewan, A3 Health Sciences Building, 107 Wiggins Road, Saskatoon, Saskatchewan, Canada S7N 5E5.