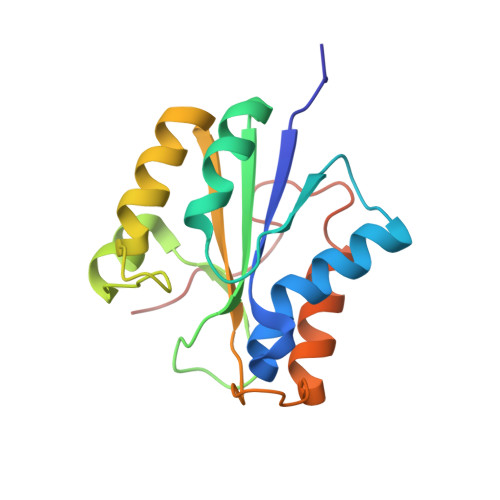
Crystal structure of hydrogenase maturating endopeptidase HycI from Escherichia coli
Kumarevel, T., Tanaka, T., Bessho, Y., Shinkai, A., Yokoyama, S.(2009) Biochem Biophys Res Commun 389: 310-314
- PubMed: 19720045
- DOI: https://doi.org/10.1016/j.bbrc.2009.08.135
- Primary Citation of Related Structures:
2E85 - PubMed Abstract:
The maturation of [NiFe]-hydrogenases is a catalyzed process involving the activities of at least seven proteins. The last step consists of the endoproteolytic cleavage of the precursor of the large subunit, after the [NiFe]-metal center has been assembled. The HycI endopeptidase is involved in the C-terminal processing of HycE, the large subunit of hydrogenase 3 from Escherichia coli. Although HycI has been well characterized biochemically, the crystallization of the protein has been quite challenging. Here, we present the crystal structure of HycI at 1.70 A resolution. The crystal structure resembles the recently reported solution structure (NMR) of the same protein and the holo-HyPD structure of the same family, but a significant conformational change is observed at the L5 loop, as compared with the solution structures of HycI and HyPD. In our crystal structure, three specific metal binding sites (Ca1-3) were identified and these metal ions are possibly involved in the C-terminal cleavage of HycE.
Organizational Affiliation:
RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo, Hyogo 679-5148, Japan. tskvel@spring8.or.jp