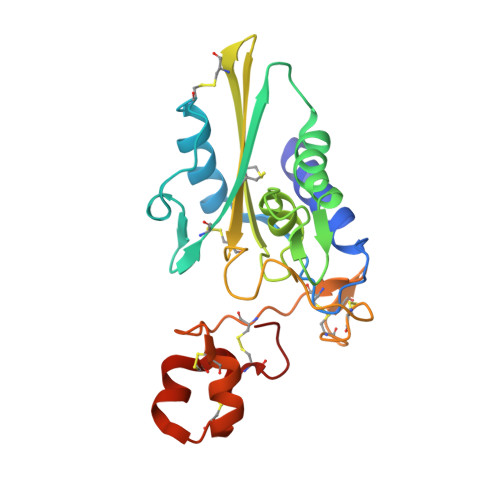
Structures of pseudechetoxin and pseudecin, two snake-venom cysteine-rich secretory proteins that target cyclic nucleotide-gated ion channels: implications for movement of the C-terminal cysteine-rich domain
Suzuki, N., Yamazaki, Y., Brown, R.L., Fujimoto, Z., Morita, T., Mizuno, H.(2008) Acta Crystallogr D Biol Crystallogr 64: 1034-1042
- PubMed: 18931410
- DOI: https://doi.org/10.1107/S0907444908023512
- Primary Citation of Related Structures:
2DDA, 2DDB, 2EPF - PubMed Abstract:
Cyclic nucleotide-gated (CNG) ion channels play pivotal roles in sensory transduction by retinal photoreceptors and olfactory neurons. The elapid snake toxins pseudechetoxin (PsTx) and pseudecin (Pdc) are the only known protein blockers of CNG channels. These toxins belong to a cysteine-rich secretory protein (CRISP) family containing an N-terminal pathogenesis-related proteins of group 1 (PR-1) domain and a C-terminal cysteine-rich domain (CRD). PsTx and Pdc are highly homologous proteins, but their blocking affinities on CNG channels are different: PsTx blocks both the olfactory and retinal channels with approximately 15-30-fold higher affinity than Pdc. To gain further insights into their structure and function, the crystal structures of PsTx, Pdc and Zn2+-bound Pdc were determined. The structures revealed that most of the amino-acid-residue differences between PsTx and Pdc are located around the concave surface formed between the PR-1 domain and the CRD, suggesting that the concave surface is functionally important for CNG-channel binding and inhibition. A structural comparison in the presence and absence of Zn2+ ion demonstrated that the concave surface can open and close owing to movement of the CRD upon Zn2+ binding. The data suggest that PsTx and Pdc occlude the pore entrance and that the dynamic motion of the concave surface facilitates interaction with the CNG channels.
Organizational Affiliation:
Department of Applied Biochemistry, University of Tsukuba, Tsukuba, Ibaraki 305-8572, Japan.