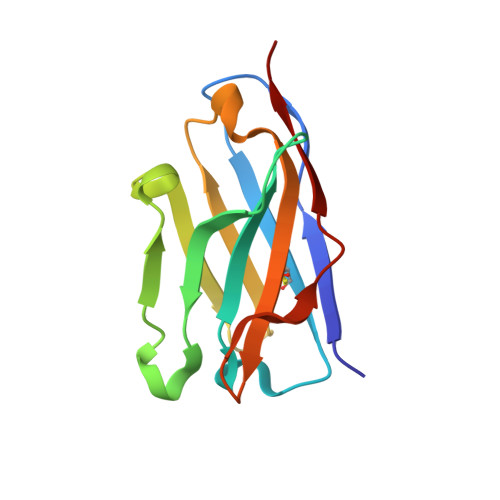
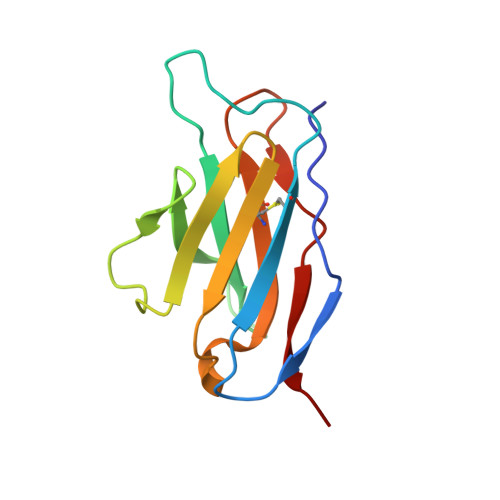
Structural Basis of Affinity Maturation of the Tepc15/V(Kappa)45.1 Anti-2-Phenyl-5-Oxazolone Antibodies.
Scotti, C., Gherardi, E.(2006) J Mol Biol 359: 1161
- PubMed: 16682055
- DOI: https://doi.org/10.1016/j.jmb.2006.04.036
- Primary Citation of Related Structures:
2CJU, 2UUD - PubMed Abstract:
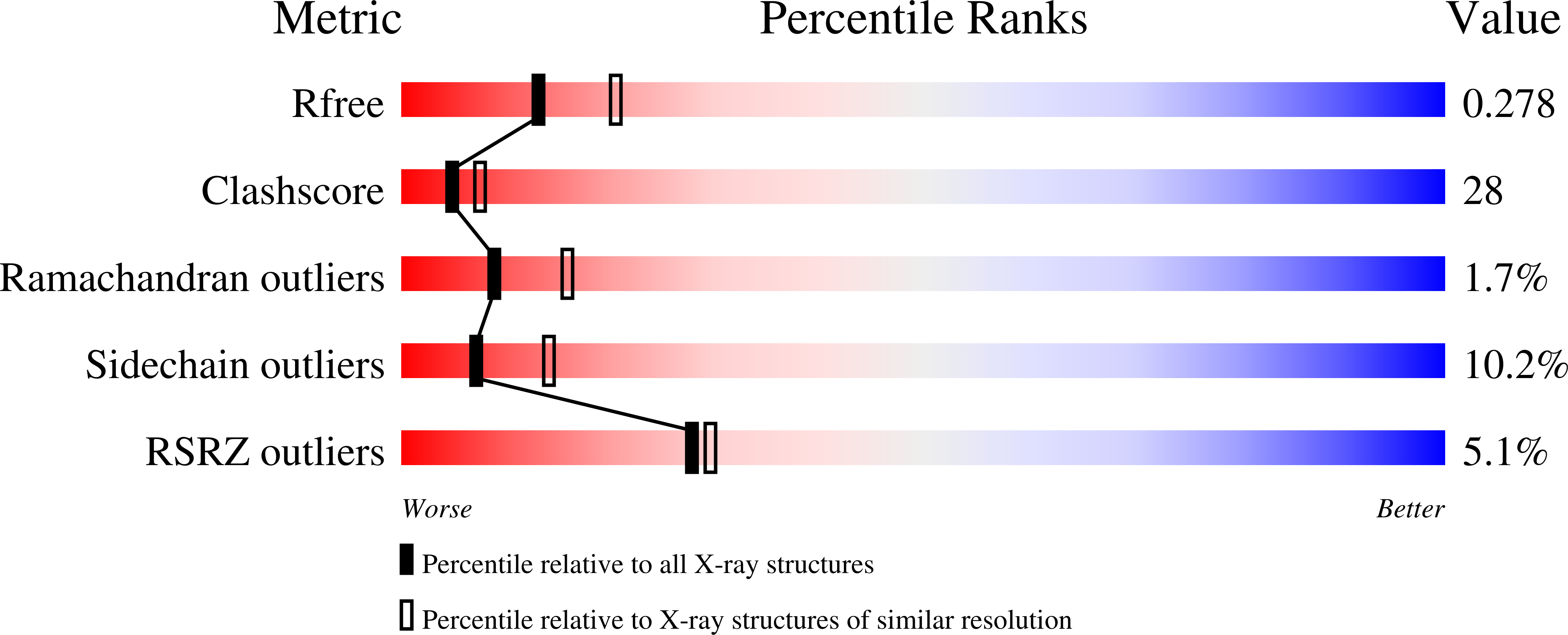
Affinity maturation is a process that leads to the emergence of more efficient antibodies following initial antigen encounter and represents a key strategy of the adaptive immunity of vertebrate organisms. Earlier and detailed sequence studies of the antibody response to a model antigen, the hapten 2-phenyl-5-oxazolone (phOx), define three different classes of antibodies. Class I antibodies use the V(H)Ox1/V(kappa)Ox1 gene pair and dominate the early stages of the anti-phOx response, class II antibodies use the V(kappa)Ox1 gene but a different V(H) segment and are common in the intermediate stages, and class III antibodies use the TEPC15/V(kappa)45.1 genes and play the greatest role in the late stages. Only the crystal structure of one anti-phOx antibody, the class II NQ10/12.5 Fab fragment, has been described. Here we report the crystal structures of the scFv form of the low and high affinity anti-phOx class III antibodies NQ10/1.12 and NQ16/113.8 complexed with the hapten. The two antibodies differ by nine amino acid substitutions, all located in the V(H) domain. Analysis of the two structures shows that affinity maturation results from an increase in surface complementarity, as a consequence of a finely tuned and highly concerted process chaperoned by the somatic mutations, and implies a more efficient hapten-induced fit in the mature antibody. The data also demonstrate that class III antibodies respond in a completely different way to the architectural problem of binding phOx compared to the class II antibody NQ10/12.5.
Organizational Affiliation:
Dipartimento di Medicina Sperimentale, Universita' di Pavia, Italy. claudia.scotti@unipv.it