Analysis of protein solvent interactions in glucose dehydrogenase from the extreme halophile Haloferax mediterranei.
Britton, K.L., Baker, P.J., Fisher, M., Ruzheinikov, S., Gilmour, D.J., Bonete, M.-J., Ferrer, J., Pire, C., Esclapez, J., Rice, D.W.(2006) Proc Natl Acad Sci U S A 103: 4846-4851
- PubMed: 16551747
- DOI: https://doi.org/10.1073/pnas.0508854103
- Primary Citation of Related Structures:
2B5V, 2B5W - PubMed Abstract:
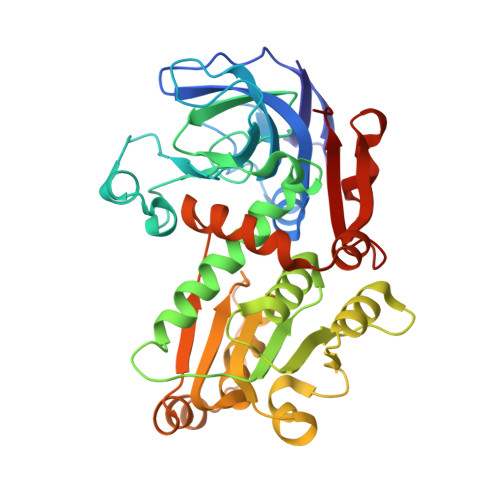
The structure of glucose dehydrogenase from the extreme halophile Haloferax mediterranei has been solved at 1.6-A resolution under crystallization conditions which closely mimic the "in vivo" intracellular environment. The decoration of the enzyme's surface with acidic residues is only partially neutralized by bound potassium counterions, which also appear to play a role in substrate binding. The surface shows the expected reduction in hydrophobic character, surprisingly not from changes associated with the loss of exposed hydrophobic residues but rather arising from a loss of lysines consistent with the genome wide-reduction of this residue in extreme halophiles. The structure reveals a highly ordered, multilayered solvation shell that can be seen to be organized into one dominant network covering much of the exposed surface accessible area to an extent not seen in almost any other protein structure solved. This finding is consistent with the requirement of the enzyme to form a protective shell in a dehydrating environment.
Organizational Affiliation:
Krebs Institute for Biomolecular Research, Department of Molecular Biology and Biotechnology, University of Sheffield, Sheffield S10 2TN, United Kingdom.