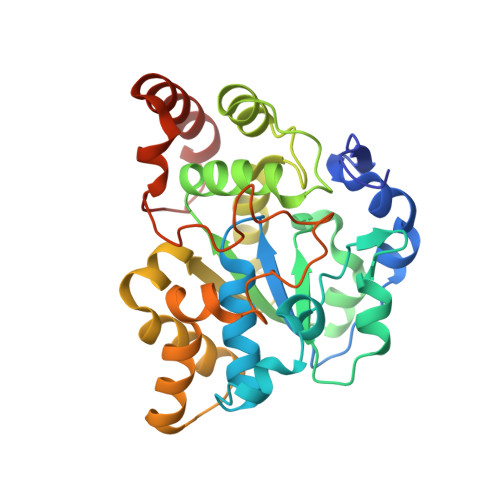
Crystal structure of mSULT1D1, a mouse catecholamine sulfotransferase
Teramoto, T., Sakakibara, Y., Inada, K., Kurogi, K., Liu, M.-C., Suiko, M., Kimura, M., Kakuta, Y.(2008) FEBS Lett 582: 3909-3914
- PubMed: 18977225
- DOI: https://doi.org/10.1016/j.febslet.2008.10.035
- Primary Citation of Related Structures:
2ZPT - PubMed Abstract:
In mammals, sulfonation as mediated by specific cytosolic sulfotransferases (SULTs) plays an important role in the homeostasis of dopamine and other catecholamines. To gain insight into the structural basis for dopamine recognition/binding, we determined the crystal structure of a mouse dopamine-sulfating SULT, mouse SULT1D1 (mSULT1D1). Data obtained indicated that mSULT1D1 comprises of a single alpha/beta domain with a five-stranded parallel beta-sheet. In contrast to the structure of the human SULT1A3 (hSULT1A3)-dopamine complex previously reported, molecular modeling and mutational analysis revealed that a water molecule plays a critical role in the recognition of the amine group of dopamine by mSULT1D1. These results imply differences in substrate binding between dopamine-sulfating SULTs from different species.
Organizational Affiliation:
Laboratory of Structural Biology, Department of Systems Life Sciences, Japan.