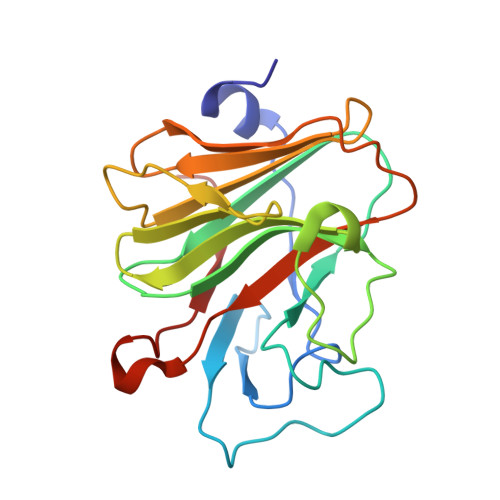
The Crystal Structure of Human Pyrin B30.2 Domain: Implications for Mutations Associated with Familial Mediterranean Fever.
Weinert, C., Gruetter, C., Roschitzki-Voser, H., Mittl, P.R., Gruetter, M.G.(2009) J Mol Biol 394: 226
- PubMed: 19729025
- DOI: https://doi.org/10.1016/j.jmb.2009.08.059
- Primary Citation of Related Structures:
2WL1 - PubMed Abstract:
The inherited autoinflammatory syndrome familial Mediterranean fever (FMF) is characterized by recurrent episodes of fever, which are independent of any bacterial or viral infections. This disease is associated with point mutations in the mefv gene product pyrin. Although the precise molecular functions of pyrin are unknown, it seems to be involved in the maturation and secretion of interleukin-1beta. Approximately two thirds of all FMF-associated mutations cluster in the C-terminal B30.2 domain of pyrin. To investigate the molecular consequences of FMF-associated mutations, we determined the crystal structure of the pyrin B30.2 domain at 1.35-A resolution. The comparison with other B30.2/ligand complex structures revealed a shallow cavity, which seems to be involved in binding the pyrin ligand. The bottom of this cavity is covered mainly with hydrophobic amino acids, suggesting that pyrin recognizes its ligand by hydrophobic contacts and surface complementarities. FMF-associated mutations cluster around two sites on the B30.2 surface. Approximately two thirds, including those mutations with the most severe disease outcomes, are observed in the vicinity of the predicted peptide binding site, suggesting that they will have a direct impact on ligand binding. A second mutational hot spot was observed on the opposite side of the B30.2 domain in the neighbourhood of its artificial N-terminus. Although most FMF-associated mutations are solvent exposed, several will modify the main-chain conformation of loops. The experimental crystal structure of the pyrin B30.2 domain serves as a basis for an accurate modelling of these mutations.
Organizational Affiliation:
Department for Biochemistry, University of Zürich, Winterthurer Str. 190, CH-8057 Zürich, Switzerland.