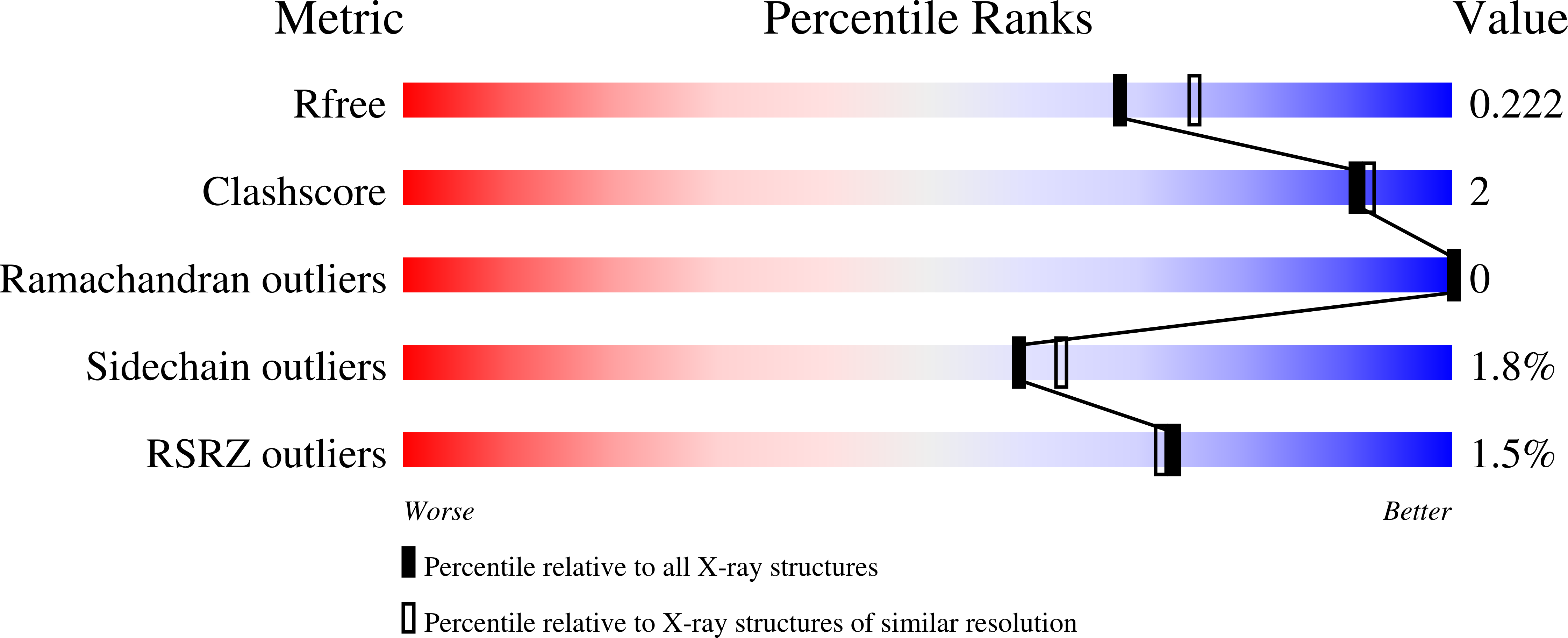
The Crystal Structure of an Lll-Configured Depsipeptide Substrate Analogue Bound to Isopenicillin N Synthase.
Ge, W., Clifton, I.J., Stok, J.E., Adlington, R.M., Baldwin, J.E., Rutledge, P.J.(2010) Org Biomol Chem 8: 122
- PubMed: 20024142
- DOI: https://doi.org/10.1039/b910170e
- Primary Citation of Related Structures:
2VBD - PubMed Abstract:
Isopenicillin N synthase (IPNS) is a non-heme iron(ii) oxidase, which catalyses the biosynthesis of isopenicillin N (IPN) from the tripeptide delta-l-alpha-aminoadipoyl-l-cysteinyl-d-valine (lld-ACV) in a remarkable oxidative bicyclisation reaction. The natural substrate for IPNS is the lld-configured tripeptide. lll-ACV is not turned over by the enzyme, but inhibits turnover of the lld-tripeptide. The mechanism by which this inhibition takes place is not fully understood. Recent studies have employed a range of lld-configured depsipeptide substrate analogues in crystallographic studies to probe events preceding beta-lactam closure in the IPNS reaction cycle. Herein, we report the first crystal structure of IPNS in complex with an lll-configured depsipeptide analogue, delta-l-alpha-aminoadipoyl-l-cysteine (1-(R)-carboxy-2-thiomethyl)ethyl ester (lll-ACOmC). This report describes the crystal structure of the IPNS:Fe(ii):lll-ACOmC complex to 2.0 A resolution, and discusses attempts to oxygenate this complex at high pressure in order to probe the mechanism by which lll-configured substrates inhibit IPNS catalysis.
Organizational Affiliation:
Chemistry Research Laboratory, University of Oxford, UK.