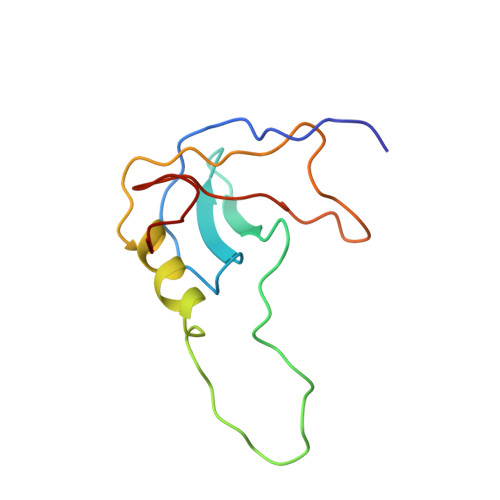
NMR structure of a stable "OB-fold" sub-domain isolated from staphylococcal nuclease.
Alexandrescu, A.T., Gittis, A.G., Abeygunawardana, C., Shortle, D.(1995) J Mol Biol 250: 134-143
- PubMed: 7608966
- DOI: https://doi.org/10.1006/jmbi.1995.0365
- Primary Citation of Related Structures:
2SOB - PubMed Abstract:
Similar folds often occur in proteins with dissimilar sequences. The OB-fold forms a part of the structures of at least seven non-homologous proteins that share either oligonucleotide or oligosaccharide binding functions. A 1-103 fragment corresponding to the OB-fold of the 149 amino acid residue staphylococcal nuclease gives NMR spectra characteristic of an unfolded protein, i.e. the wild-type nuclease sequence is insufficient to maintain a stable tertiary structure in the absence of the C-terminal one-third of this single-domain protein. By contrast, the 1-103 fragment of nuclease with the mutations Val66Leu and Gly88Val adopts a stable tertiary structure. The NMR solution structure of this latter fragment is a close variation of the OB-fold found in the X-ray structure of the parent protein. The Val66Leu and Gly88Val mutations appear to stabilize tertiary structure by consolidating the hydrophobic core of the nuclease OB-fold sub-domain. Taken together, these results suggest that recurrent structural motifs such as the OB-fold may in some cases represent vestiges of autonomous folding units that, during evolution, have become integrated into more complex cooperative folding domains.
Organizational Affiliation:
Department of Biological Chemistry, Johns Hopkins Medical Institutions, Baltimore, MD 21205, USA.