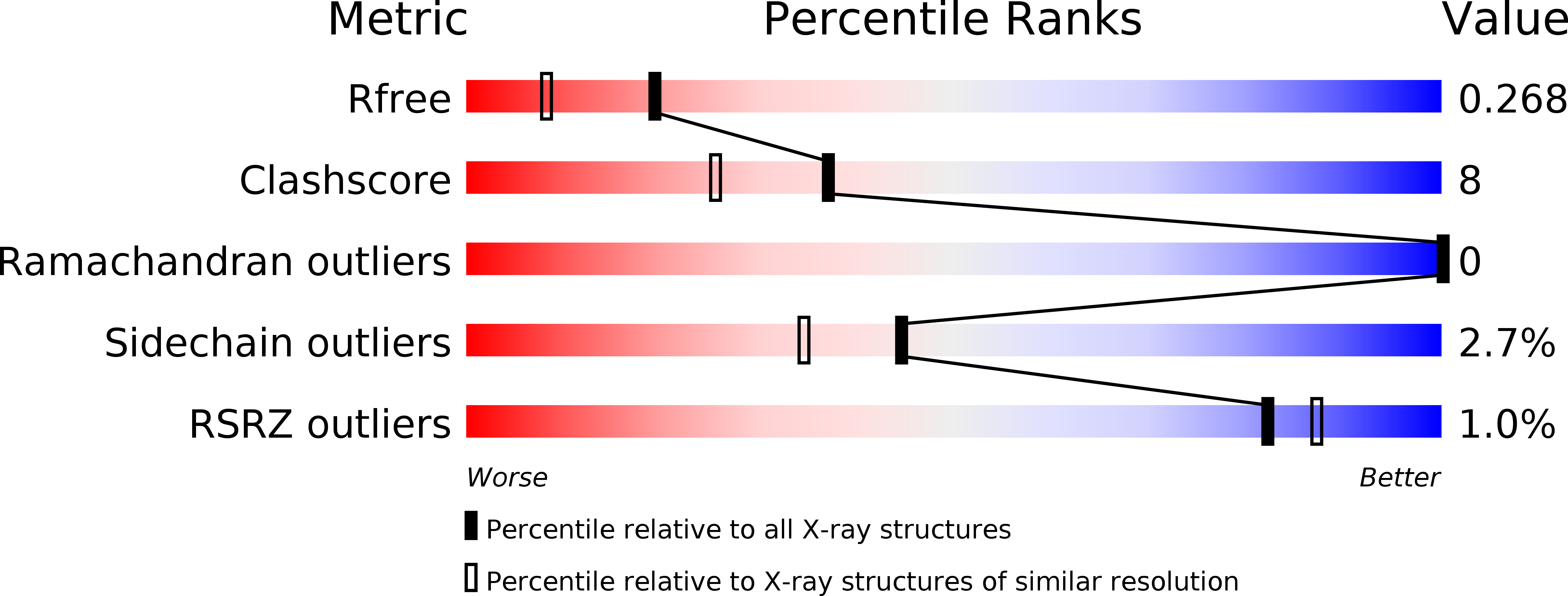
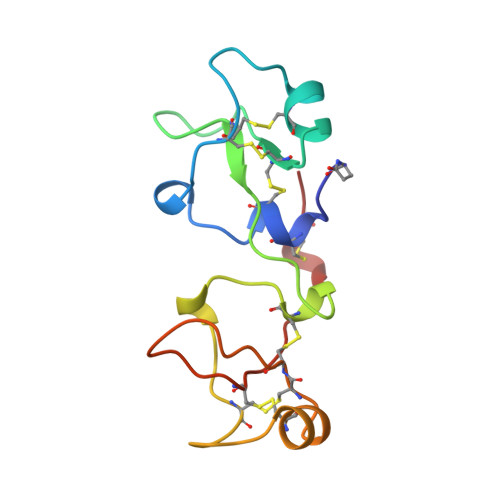
Structure of porcine pancreatic spasmolytic polypeptide at 1.95 A resolution.
Petersen, T.N., Henriksen, A., Gajhede, M.(1996) Acta Crystallogr D Biol Crystallogr 52: 730-737
- PubMed: 15299636
- DOI: https://doi.org/10.1107/S0907444996001345
- Primary Citation of Related Structures:
2PSP - PubMed Abstract:
The structure of a trigonal crystal form of porcine pancreatic spasmolytic polypeptide (PSP) has been solved by molecular replacement and refined to 1.95 A resolution. Three heavy-atom derivatives were prepared, giving unbiased phase information, which was used in the model building of the protein molecules. The final conventional R value is 19.8% with the inclusion of 183 water molecules. PSP crystallizes as a dimer in space group P3(1)21 with a non-crystallographic twofold axis relating the monomers. The monomer consists of two very similar domains each composed of three loop regions. Two clefts are found in the monomer, one in each domain, that are proposed as possible substrate-binding sites. Important interactions have been identified in the proposed substrate-binding sites, where conserved water molecules probably mimic the hydrophilic positions of the substrate. The estimated cleft size is 9 x 9 x 12 A. Analysis of the charge distribution within the clefts, by an electrostatic potential calculation, shows the clefts to be essentially non-charged.
Organizational Affiliation:
Centre for Crystallographic Studies, Universitetsparken, Department of Chemsistry, University of Copenhagen, Denmark.