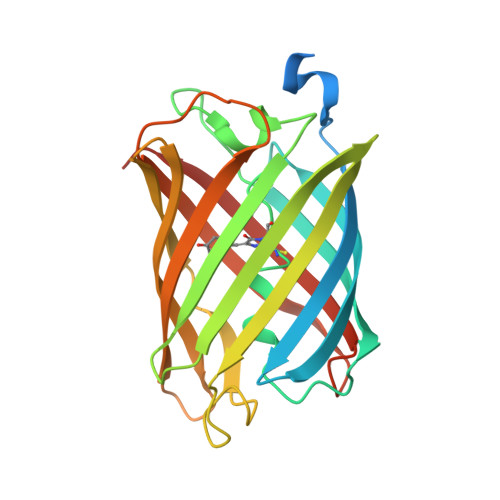
Structural basis for reversible photoswitching in Dronpa
Andresen, M., Stiel, A.C., Trowitzsch, S., Weber, G., Eggeling, C., Wahl, M.C., Hell, S.W., Jakobs, S.(2007) Proc Natl Acad Sci U S A 104: 13005-13009
- PubMed: 17646653
- DOI: https://doi.org/10.1073/pnas.0700629104
- Primary Citation of Related Structures:
2POX - PubMed Abstract:
Dronpa is a novel GFP-like fluorescent protein with exceptional light-controlled switching properties. It may be reversibly switched between a fluorescent on-state and a nonfluorescent off-state by irradiation with light. To elucidate the molecular basis of the switching mechanism, we generated reversibly switchable Dronpa protein crystals. Using these crystals we determined the elusive dark-state structure of Dronpa at 1.95-A resolution. We found that the photoswitching results in a cis-trans isomerization of the chromophore accompanied by complex structural rearrangements of four nearby amino acid residues. Because of this cascade of intramolecular events, the chromophore is exposed to distinct electrostatic surface potentials, which are likely to influence the protonation equilibria at the chromophore. We suggest a comprehensive model for the light-induced switching mechanism, connecting a cascade of structural rearrangements with different protonation states of the chromophore.
Organizational Affiliation:
Department of NanoBiophotonics, Max Planck Institute for Biophysical Chemistry, Am Fassberg 11, 37077 Göttingen, Germany.