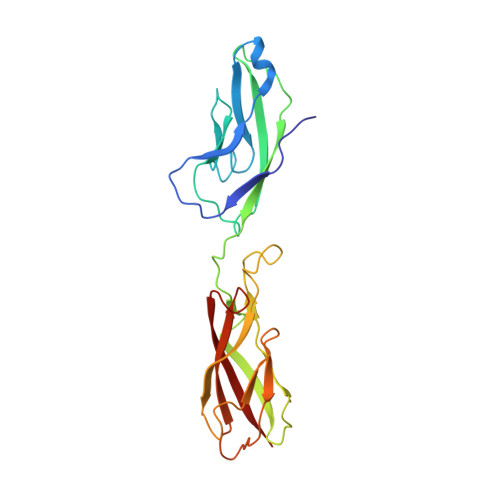
The Crystal Structure of Human E-cadherin Domains 1 and 2, and Comparison with other Cadherins in the Context of Adhesion Mechanism
Parisini, E., Higgins, J.M.G., Liu, J.-H., Brenner, M.B., Wang, J.-H.(2007) J Mol Biol 373: 401-411
- PubMed: 17850815
- DOI: https://doi.org/10.1016/j.jmb.2007.08.011
- Primary Citation of Related Structures:
2O72 - PubMed Abstract:
Cell adhesion mediated by type I cadherins involves homophilic "trans" interactions that are thought to be brought about by a strand exchange mechanism involving the N-terminal extracellular domain. Here, we present the high-resolution crystal structure of the N-terminal two domains of human E-cadherin. Comparison of this structure with other type I cadherin structures reveals features that are likely to be critical to facilitate dimerization by strand exchange as well as dimer flexibility. We integrate this structural knowledge to provide a model for type I cadherin adhesive interactions. Intra-molecular docking of the conserved N-terminal "adhesion arm" into the acceptor pocket in monomeric E-cadherin appears largely identical to inter-molecular docking of the adhesion arm in adhesive trans dimers. A strained conformation of the adhesion arm in the monomer, however, may create an equilibrium between "open" and "closed" forms that primes the cadherin for formation of adhesive interactions, which are then stabilized by additional dimer-specific contacts. By contrast, in type II cadherins, strain in the adhesion arm appears absent and a much larger surface area is involved in trans adhesion, which may compensate the activation energy required to peel off the intra-molecularly docked arm. It seems that evolution has selected slightly different adhesion mechanisms for type I and type II cadherins.
Organizational Affiliation:
Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA 02115, USA.