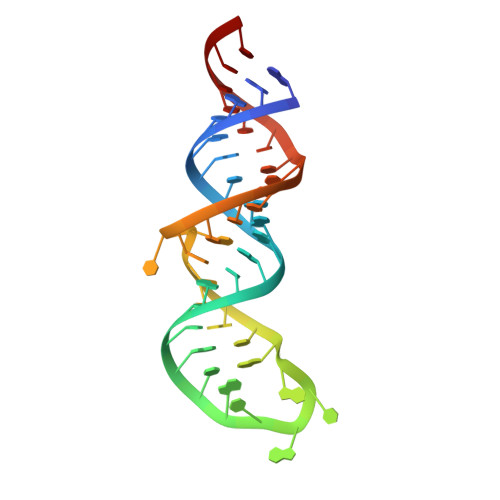
Preformed protein-binding motifs in 7SK snRNA: structural and thermodynamic comparisons with retroviral TAR.
Durney, M.A., D'Souza, V.M.(2010) J Mol Biol 404: 555-567
- PubMed: 20816986
- DOI: https://doi.org/10.1016/j.jmb.2010.08.042
- Primary Citation of Related Structures:
2KX8 - PubMed Abstract:
The 7SK small nuclear RNA is a highly conserved non-coding RNA that regulates transcriptional elongation. 7SK utilizes the HEXIM proteins to sequester the transcription factor P-TEFb by a mechanism similar to that used by retroviral TAR RNA to engage Tat and P-TEFb. Tat has also recently been shown to bind 7SK directly and recruit P-TEFb to TAR. We report here the solution structures of the free and arginine-bound forms of stem loop 4 of 7SK (7SK-SL4). Comparison of the 7SK-SL4 and TAR structures demonstrates the presence of a common arginine sandwich motif. However, arginine binding to 7SK-SL4 is mechanistically distinct and occurs via docking into a pre-organized pocket resulting in a 1000-fold increased affinity. Furthermore, whereas formation of the binding pocket in TAR requires a critical base-triple, hydrogen-bond formation between the equivalent bases in 7SK-SL4 is not essential and the pocket is stabilized solely by a pseudo base-triple platform. In addition, this theme of preformed protein binding motifs also extends into the pentaloop. The configuration of the loop suggests that 7SK-SL4 is poised to make ternary contacts with P-TEFb and HEXIM or Tat. These key differences between 7SK-SL4 and TAR present an opportunity to understand RNA structural adaptation and have implications for understanding differential interactions with Tat.
Organizational Affiliation:
Department of Molecular and Cellular Biology, Harvard University, Cambridge, MA 02138, USA.