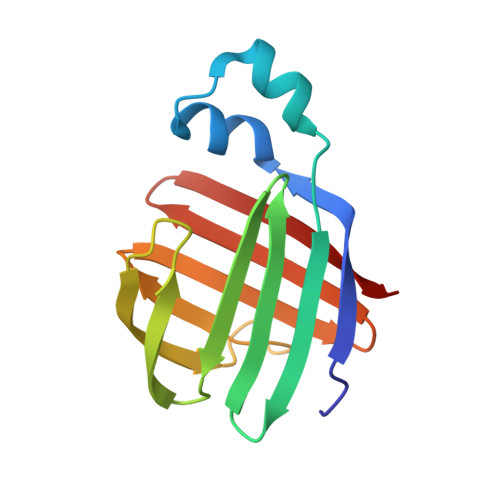
Solution-State Molecular Structure of Apo and Oleate-Liganded Liver Fatty Acid-Binding Protein
He, Y., Yang, X., Wang, H., Estephan, R., Francis, F., Kodukula, S., Storch, J., Stark, R.E.(2007) Biochemistry 46: 12543-12556
- PubMed: 17927211
- DOI: https://doi.org/10.1021/bi701092r
- Primary Citation of Related Structures:
2JU3, 2JU7, 2JU8 - PubMed Abstract:
Rat liver fatty acid-binding protein (LFABP) is distinctive among intracellular lipid-binding proteins (iLBPs): more than one molecule of long-chain fatty acid and a variety of diverse ligands can be bound within its large cavity, and in vitro lipid transfer to model membranes follows a mechanism that is diffusion-controlled rather than mediated by protein-membrane collisions. Because the apoprotein has proven resistant to crystallization, nuclear magnetic resonance spectroscopy offers a unique route to functionally informative comparisons of molecular structure and dynamics for LFABP in free (apo) and liganded (holo) forms. We report herein the solution-state structures determined for apo-LFABP at pH 6.0 and for holoprotein liganded to two oleates at pH 7.0, as well as the structure of the complex including locations of the ligands. 1H, 13C, and 15N resonance assignments revealed very similar types and locations of secondary structural elements for apo- and holo-LFABP as judged from chemical shift indices. The solution-state tertiary structures of the proteins were derived with the CNS/ARIA computational protocol, using distance and angular restraints based on 1H-1H nuclear Overhauser effects (NOEs), hydrogen-bonding networks, 3J(HNHA) coupling constants, intermolecular NOEs, and residual dipolar (NH) couplings. The holo-LFABP solution-state conformation is in substantial agreement with a previously reported X-ray structure [Thompson, J., Winter, N., Terwey, D., Bratt, J., and Banaszak, L. (1997) The crystal structure of the liver fatty acid-binding protein. A complex with two bound oleates, J. Biol. Chem. 272, 7140-7150], including the typical beta-barrel capped by a helix-turn-helix portal. In the solution state, the internally bound oleate has the expected U-shaped conformation and is tethered electrostatically, but the extended portal ligand can adopt a range of conformations based on the computationally refined structures, in contrast to the single conformation observed in the crystal structure. The apo-LFABP also has a well-defined beta-barrel structural motif typical of other members of the iLBP protein family, but the portal region that is thought to facilitate ligand entry and exit exhibits conformational variability and an unusual "open cap" orientation with respect to the barrel. These structural results allow us to propose a model in which ligand binding to LFABP occurs through conformational fluctuations that adjust the helix-turn-helix motif to open or close the top of the beta-barrel, and solvent accessibility to the protein cavity favors diffusion-controlled ligand transport.
Organizational Affiliation:
Department of Chemistry, College of Staten Island, and Graduate Center and Institute for Macromolecular Assemblies, City University of New York, 2800 Victory Boulevard, Staten Island, New York 10314-6600, USA.