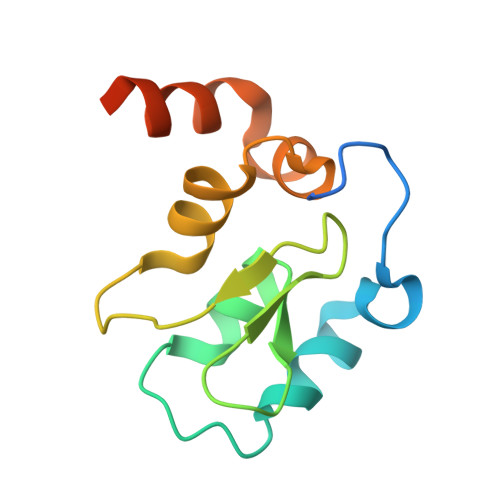
Structure-Based Design, Synthesis, Evaluation, and Crystallographic Studies of Conformationally Constrained Smac Mimetics as Inhibitors of the X-Linked Inhibitor of Apoptosis Protein (Xiap).
Sun, H., Stuckey, J.A., Nikolovska-Coleska, Z., Qin, D., Meagher, J.L., Qiu, S., Lu, J., Yang, C., Saito, N.G., Wang, S.(2008) J Med Chem 51: 7169
- PubMed: 18954041
- DOI: https://doi.org/10.1021/jm8006849
- Primary Citation of Related Structures:
2JK7 - PubMed Abstract:
Small molecules designed to mimic the binding of Smac protein to X-linked inhibitor of apoptosis protein (XIAP) are being pursued as a promising new class of anticancer drugs. Herein, we report the design, synthesis, and comprehensive structure-activity relationship studies of a series of conformationally constrained bicyclic Smac mimetics. Our studies led to the discovery of a number of highly potent and cell-permeable Smac mimetics and yielded important new insights into their structure-activity relationship for their binding to XIAP and for their activity in inhibition of cancer cell growth. Determination of the crystal structure of one potent Smac mimetic, compound 21, in complex with XIAP BIR3 provides the structural basis for its high-affinity binding to XIAP and for the design of highly potent Smac mimetics.
Organizational Affiliation:
Department of Internal Medicine, University of Michigan, Ann Arbor, Michigan 48109, USA.