New Insights on the Role of the Gamma-Herpesvirus Uracil-DNA Glycosylase Leucine Loop Revealed by the Structure of the Epstein-Barr Virus Enzyme in Complex with an Inhibitor Protein.
Geoui, T., Buisson, M., Tarbouriech, N., Burmeister, W.P.(2007) J Mol Biol 366: 117
- PubMed: 17157317
- DOI: https://doi.org/10.1016/j.jmb.2006.11.007
- Primary Citation of Related Structures:
2J8X - PubMed Abstract:
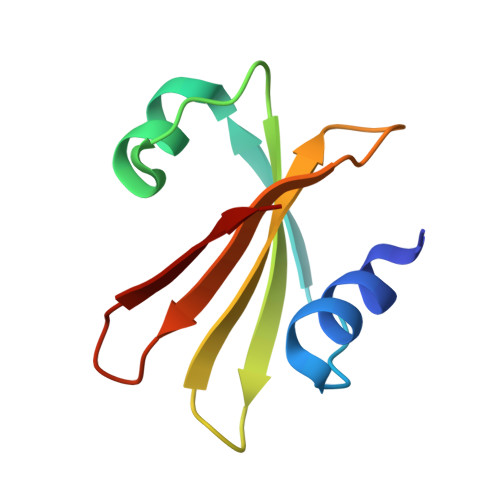
Epstein-Barr virus (EBV) is a human gamma-herpesvirus. Within its 86 open reading frame containing genome, two enzymes avoiding uracil incorporation into DNA can be found: uracil triphosphate hydrolase and uracil-DNA glycosylase (UNG). The latter one excises uracil bases that are due to cytosine deamination or uracil misincorporation from double-stranded DNA substrates. The EBV enzyme belongs to family 1 UNGs. We solved the three-dimensional structure of EBV UNG in complex with the uracil-DNA glycosylase inhibitor protein (Ugi) from bacteriophage PBS-2 at a resolution of 2.3 A by X-ray crystallography. The structure of EBV UNG encoded by the BKRF3 reading frame shows the excellent global structural conservation within the solved examples of family 1 enzymes. Four out of the five catalytic motifs are completely conserved, whereas the fifth one, the leucine loop, carries a seven residue insertion. Despite this insertion, catalytic constants of EBV UNG are similar to those of other UNGs. Modelling of the EBV UNG-DNA complex shows that the longer leucine loop still contacts DNA and is likely to fulfil its role of DNA binding and deformation differently than the enzymes with previously solved structures. We could show that despite the evolutionary distance of EBV UNG from the natural host protein, bacteriophage Ugi binds with an inhibitory constant of 8 nM to UNG. This is due to an excellent specificity of Ugi for conserved elements of UNG, four of them corresponding to catalytic motifs and a fifth one corresponding to an important beta-turn structuring the catalytic site.
Organizational Affiliation:
Institut de Virologie Moléculaire et Structurale, FRE 2854 CNRS-UJF, BP 181, F-38042 Grenoble cedex 9, France.