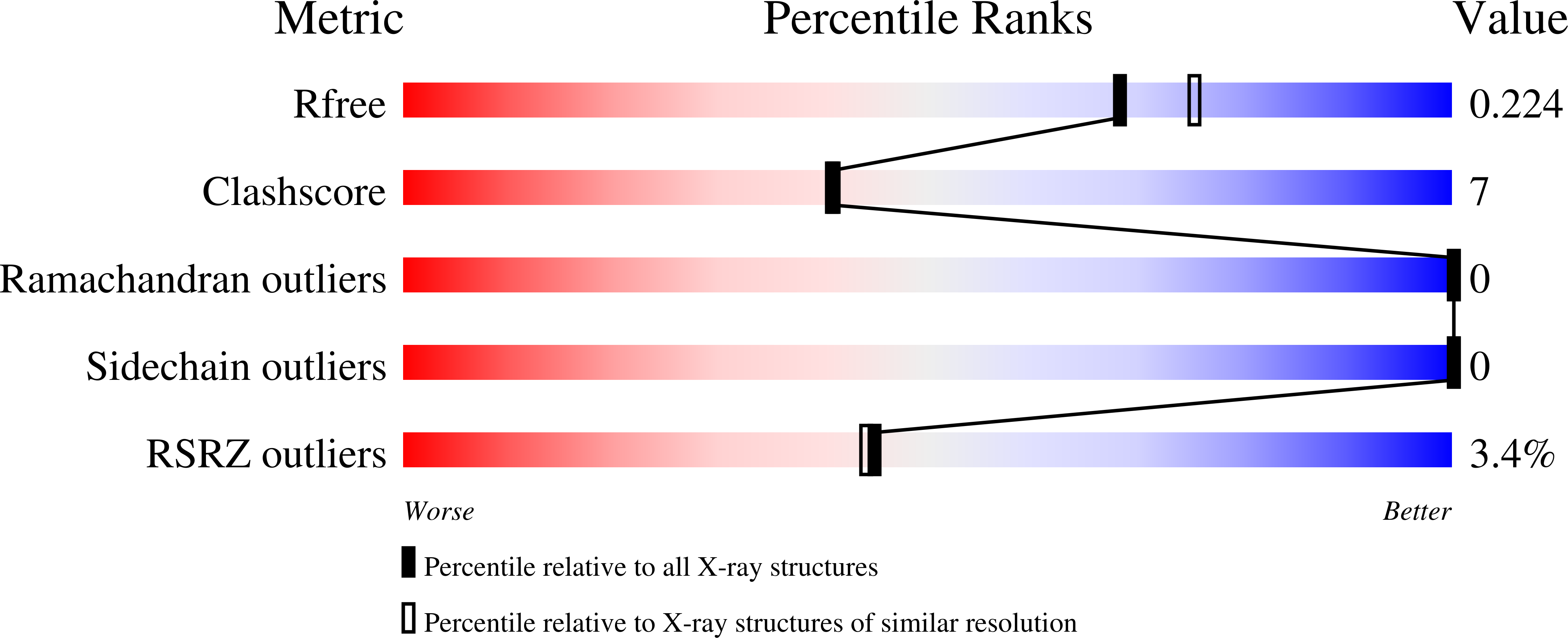
Kinetic and structural studies of specific protein-protein interactions in substrate catalysis by Cdc25B phosphatase.
Sohn, J., Buhrman, G., Rudolph, J.(2007) Biochemistry 46: 807-818
- PubMed: 17223702
- DOI: https://doi.org/10.1021/bi061257y
- Primary Citation of Related Structures:
2IFD, 2IFV - PubMed Abstract:
Using a combination of steady-state and single-turnover kinetics, we probe substrate association, dissociation, and chemistry for the reaction of Cdc25B phosphatase with its Cdk2-pTpY/CycA protein substrate. The rate constant for substrate association for the wild-type enzyme is 1.3 x 10(6) M(-1) s(-1). The rate constant for dissociation is slow compared to the rate constant for phosphate transfer to form the phospho-enzyme intermediate (k2 = 1.1 s(-1)), making Cdk2-pTpY/CycA a sticky substrate. Compared to the wild type, all hotspot mutants of residues at the remote docking site that specifically affect catalysis with the protein substrate (Arg488, Arg492, and Tyr497 on Cdc25B and Asp206 on Cdk2) have greatly slowed rate constants of association (70- to 4500-fold), and some mutants have decreased k2 values compared to that of the wild type. Most dramatically, R492L, despite showing no significant changes in a crystal structure at 2.0 A resolution, has an approximately 100-fold decrease in k2 compared to that of wild-type Cdc25B. The active site C473S mutant binds tightly to and dissociates slowly from Cdk2-pTpY/CycA (Kd = 10 nM, k(off) = 0.01 s(-1)). In contrast, the C473D mutant, despite showing only localized perturbations in the active site at 1.6 A resolution, has a much weaker affinity and dissociates rapidly (Kd of 2 microM, k(off) > 2 s(-1)) from the protein substrate. Overall, we demonstrate that the association of Cdc25B with its Cdk2-pTpY/CycA substrate is governed to a significant extent by the interactions of the remote hotspot residues, whereas dissociation is governed by interactions at the active site.
Organizational Affiliation:
Department of Biochemistry, Duke University Medical Center, Durham, North Carolina 27710, USA.