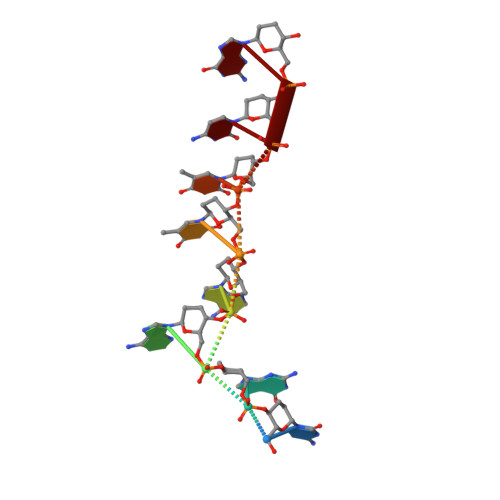
Crystal structure of homo-DNA and nature's choice of pentose over hexose in the genetic system.
Egli, M., Pallan, P.S., Pattanayek, R., Wilds, C.J., Lubini, P., Minasov, G., Dobler, M., Leumann, C.J., Eschenmoser, A.(2006) J Am Chem Soc 128: 10847-10856
- PubMed: 16910680
- DOI: https://doi.org/10.1021/ja062548x
- Primary Citation of Related Structures:
2H9S - PubMed Abstract:
An experimental rationalization of the structure type encountered in DNA and RNA by systematically investigating the chemical and physical properties of alternative nucleic acids has identified systems with a variety of sugar-phosphate backbones that are capable of Watson-Crick base pairing and in some cases cross-pairing with the natural nucleic acids. The earliest among the model systems tested to date, (4' --> 6')-linked oligo(2',3'-dideoxy-beta-d-glucopyranosyl)nucleotides or homo-DNA, shows stable self-pairing, but the pairing rules for the four natural bases are not the same as those in DNA. However, a complete interpretation and understanding of the properties of the hexapyranosyl (4' --> 6') family of nucleic acids has been impeded until now by the lack of detailed 3D-structural data. We have determined the crystal structure of a homo-DNA octamer. It reveals a weakly twisted right-handed duplex with a strong inclination between the hexose-phosphate backbones and base-pair axes, and highly irregular values for helical rise and twist at individual base steps. The structure allows a rationalization of the inability of allo-, altro-, and glucopyranosyl-based oligonucleotides to form stable pairing systems.
Organizational Affiliation:
Department of Biochemistry, School of Medicine, Vanderbilt University, Nashville, Tennessee 37232, USA. martin.egli@vanderbilt.edu