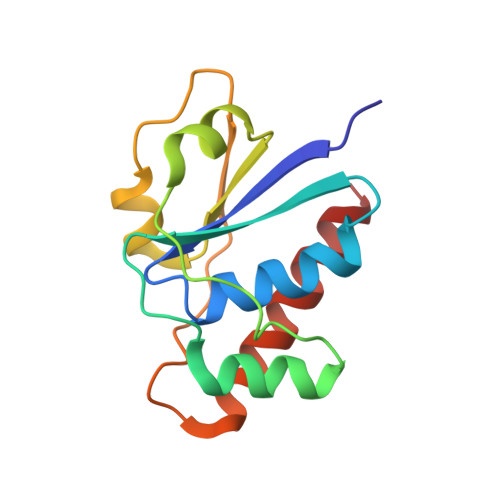
Interplay Between Ion Binding and Catalysis in the Thioredoxin-coupled Arsenate Reductase Family.
Roos, G., Buts, L., Van Belle, K., Brosens, E., Geerlings, P., Loris, R., Wyns, L., Messens, J.(2006) J Mol Biol 360: 826-838
- PubMed: 16797027
- DOI: https://doi.org/10.1016/j.jmb.2006.05.054
- Primary Citation of Related Structures:
2CD7, 2FXI - PubMed Abstract:
In the thioredoxin (Trx)-coupled arsenate reductase family, arsenate reductase from Staphylococcus aureus plasmid pI258 (Sa_ArsC) and from Bacillus subtilis (Bs_ArsC) are structurally related detoxification enzymes. Catalysis of the reduction of arsenate to arsenite involves a P-loop (Cys10Thr11Gly12Asn13Ser14Cys15Arg16) structural motif and a disulphide cascade between three conserved cysteine residues (Cys10, Cys82 and Cys89). For its activity, Sa_ArsC benefits from the binding of tetrahedral oxyanions in the P-loop active site and from the binding of potassium in a specific cation-binding site. In contrast, the steady-state kinetic parameters of Bs_ArsC are not affected by sulphate or potassium. The commonly occurring mutation of a histidine (H62), located about 6 A from the potassium-binding site in Sa_ArsC, to a glutamine uncouples the kinetic dependency on potassium. In addition, the binding affinity for potassium is affected by the presence of a lysine (K33) or an aspartic acid (D33) in combination with two negative charges (D30 and E31) on the surface of Trx-coupled arsenate reductases. In the P-loop of the Trx-coupled arsenate reductase family, the peptide bond between Gly12 and Asn13 can adopt two distinct conformations. The unique geometry of the P-loop with Asn13 in beta conformation, which is not observed in structurally related LMW PTPases, is stabilized by tetrahedral oxyanions and decreases the pK(a) value of Cys10 and Cys82. Tetrahedral oxyanions stabilize the P-loop in its catalytically most active form, which might explain the observed increase in k(cat) value for Sa_ArsC. Therefore, a subtle interplay of potassium and sulphate dictates the kinetics of Trx-coupled arsenate reductases.
Organizational Affiliation:
Laboratorium voor Ultrastructuur, Department of Molecular and Cellular Interactions, Vlaams Interuniversitair Instituut voor Biotechnologie (VIB), Vrije Universiteit Brussel (VUB), Pleinlaan 2, 1050 Brussels, Belgium.