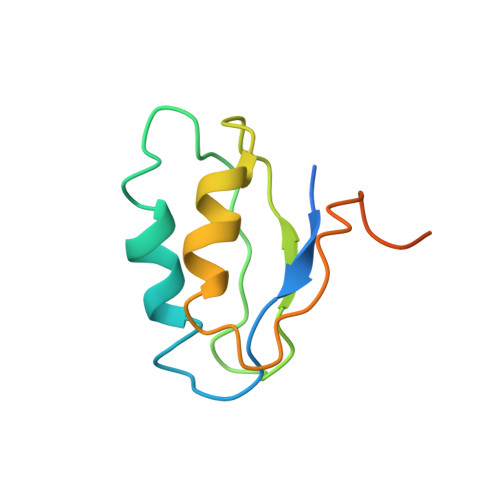
Structure determination of a new protein from backbone-centered NMR data and NMR-assisted structure prediction.
Mayer, K.L., Qu, Y., Bansal, S., LeBlond, P.D., Jenney, F.E., Brereton, P.S., Adams, M.W., Xu, Y., Prestegard, J.H.(2006) Proteins 65: 480-489
- PubMed: 16927360
- DOI: https://doi.org/10.1002/prot.21119
- Primary Citation of Related Structures:
2F40 - PubMed Abstract:
Targeting of proteins for structure determination in structural genomic programs often includes the use of threading and fold recognition methods to exclude proteins belonging to well-populated fold families, but such methods can still fail to recognize preexisting folds. The authors illustrate here a method in which limited amounts of structural data are used to improve an initial homology search and the data are subsequently used to produce a structure by data-constrained refinement of an identified structural template. The data used are primarily NMR-based residual dipolar couplings, but they also include additional chemical shift and backbone-nuclear Overhauser effect data. Using this methodology, a backbone structure was efficiently produced for a 10 kDa protein (PF1455) from Pyrococcus furiosus. Its relationship to existing structures and its probable function are discussed.
Organizational Affiliation:
Complex Carbohydrate Research Center, University of Georgia, Athens, Georgia 30602, USA.