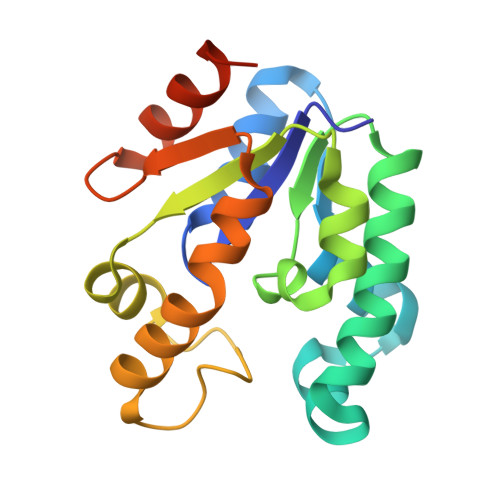
Effects of the magnesium and chloride ions and shikimate on the structure of shikimate kinase from Mycobacterium tuberculosis
Dias, M.V., Faim, L.M., Vasconcelos, I.B., de Oliveira, J.S., Basso, L.A., Santos, D.S., de Azevedo, W.F.(2007) Acta Crystallogr Sect F Struct Biol Cryst Commun 63: 1-6
- PubMed: 17183161
- DOI: https://doi.org/10.1107/S1744309106046823
- Primary Citation of Related Structures:
2DFN, 2DFT - PubMed Abstract:
Bacteria, fungi and plants can convert carbohydrate and phosphoenolpyruvate into chorismate, which is the precursor of various aromatic compounds. The seven enzymes of the shikimate pathway are responsible for this conversion. Shikimate kinase (SK) is the fifth enzyme in this pathway and converts shikimate to shikimate-3-phosphate. In this work, the conformational changes that occur on binding of shikimate, magnesium and chloride ions to SK from Mycobacterium tuberculosis (MtSK) are described. It was observed that both ions and shikimate influence the conformation of residues of the active site of MtSK. Magnesium influences the conformation of the shikimate hydroxyl groups and the position of the side chains of some of the residues of the active site. Chloride seems to influence the affinity of ADP and its position in the active site and the opening length of the LID domain. Shikimate binding causes a closing of the LID domain and also seems to influence the crystallographic packing of SK. The results shown here could be useful for understanding the catalytic mechanism of SK and the role of ions in the activity of this protein.
Organizational Affiliation:
Programa de Pós-Graduação em Biofísica Molecular, Departamento de Física, UNESP, São José do Rio Preto, SP 15054-000, Brazil.