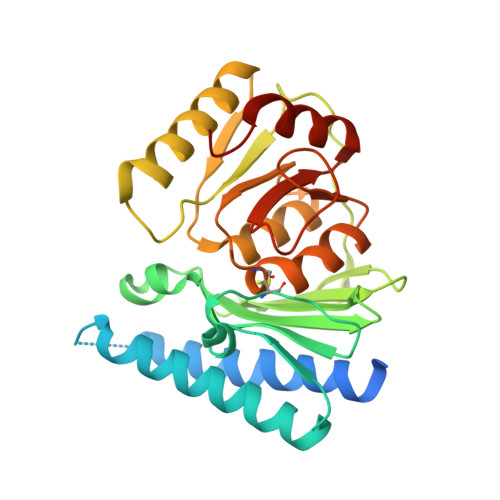
Crystal structure of human myo-inositol monophosphatase 2, the product of the putative susceptibility gene for bipolar disorder, schizophrenia, and febrile seizures
Arai, R., Ito, K., Ohnishi, T., Ohba, H., Akasaka, R., Bessho, Y., Hanawa-Suetsugu, K., Yoshikawa, T., Shirouzu, M., Yokoyama, S.(2007) Proteins 67: 732-742
- PubMed: 17340635
- DOI: https://doi.org/10.1002/prot.21299
- Primary Citation of Related Structures:
2CZH, 2CZI, 2CZK, 2DDK - PubMed Abstract:
The human IMPA2 gene, which encodes myo-inositol monophosphatase 2 (IMPA2), is mapped onto 18p11.2, a susceptibility region for bipolar disorder. This chromosomal region has also been proposed to include a susceptibility locus for schizophrenia and febrile seizures. Here we report the crystal structures of human IMPA2 and its complex with calcium and phosphate ions. Human IMPA2 comprises an alpha-beta protein with a five-layered sandwich of alpha-helices and beta-sheets (alpha-beta-alpha-beta-alpha). The crystal structure and analytical ultracentrifugation results indicated that IMPA2 exists as a dimer in solution. The overall structure of IMPA2 is similar to that of IMPA1, except for the loop regions. In IMPA1, the loop region (31-43) is located at the entrance of the active site cavity. In the corresponding region (42-54) of IMPA2, the residues are disordered and partially form an alpha-helix. The structural difference in the opening of the active site cavity suggests that the substrate specificity differs between IMPA1 and IMPA2. The widely opened cavity of IMPA2 implies that the physiological substrate may be a larger compound than inositol monophosphate. The structure of IMPA2 complexed with Ca2+ revealed two metals and one phosphate binding sites, which were the same sites as in IMPA1 complexed with Mn2+ and phosphate, suggesting that the mechanism of the enzymatic reaction is similar to that of IMPA1. The crystal structures of human IMPA2 are useful for understanding the effect of nonsynonymous polymorphism reported in IMPA2, and will contribute to further functional analyses of IMPA2 that potentially predisposes to the vulnerabilities of bipolar disorder, schizophrenia, and febrile seizures.
Organizational Affiliation:
Protein Research Group, RIKEN Genomic Sciences Center, Tsurumi, Yokohama 230-0045, Japan.