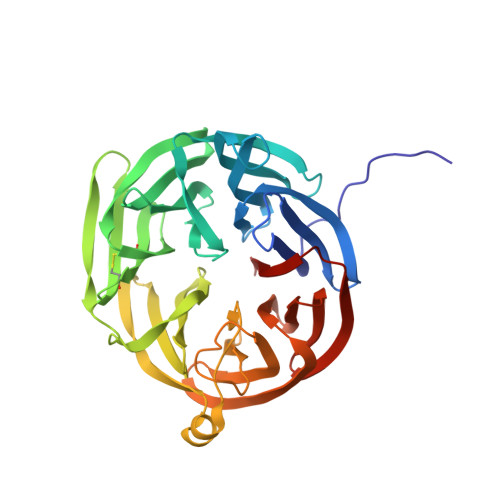
Refined crystal structure of methylamine dehydrogenase from Paracoccus denitrificans at 1.75 A resolution.
Chen, L., Doi, M., Durley, R.C., Chistoserdov, A.Y., Lidstrom, M.E., Davidson, V.L., Mathews, F.S.(1998) J Mol Biol 276: 131-149
- PubMed: 9514722
- DOI: https://doi.org/10.1006/jmbi.1997.1511
- Primary Citation of Related Structures:
2BBK - PubMed Abstract:
The three-dimensional structure of the quinoprotein methylamine dehydrogenase from Paracoccus denitrificans has been refined at 1.75 A resolution utilizing the DNA-based protein sequence. The final model incorporates 8034 atoms per molecule, including 552 molecules of solvent, and gives an R-factor of 0.163. The molecule is an H2L2 hetero-tetramer containing a non-crystallographic 2-fold axis of symmetry. The 373-residue H subunit is folded into seven repeats of a four-stranded antiparallel beta-sheet motif, arranged in a propeller-like pattern about a pseudo-7-fold rotational axis of symmetry. Each L subunit contains 131 residues folded in a tight structure composed of five beta-strands in two sheets and crosslinked by six disulfide bonds. In addition there is an intrasubunit covalent linkage between two tryptophan side-chains that form the unique redox center, tryptophan tryptophylquinone (TTQ). The active site contains the O-6 carbonyl of TTQ, the side-chains of Asp32L Asp76L, Tyr119L and Thr122L, and two solvent molecules. A potential "gate" (Phe55H) separates the closed active-site cavity from a channel containing a group of highly ordered water molecules to bulk solvent. Phe55H and Tyr119L, and a number of neighboring oxygen atoms, may also provide a binding site for monovalent cations that are known to affect the reactivity and spectral properties of TTQ as well as the oxidative half reaction. The overall reaction has been dissected into a number of discrete steps that may require participation by several individual amino acid residues in the active site acting as general acids and bases.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Washington University Medical School, St. Louis, MO 63110, USA.