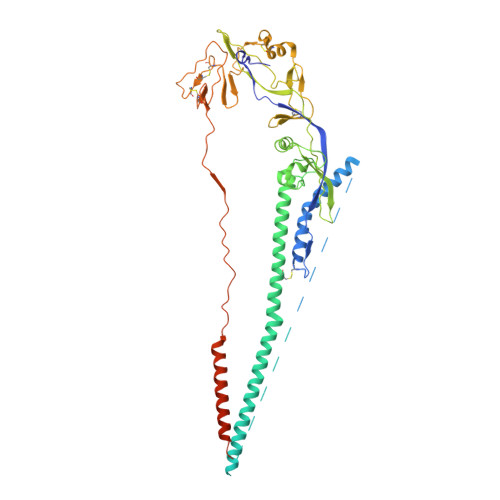
Structure of the uncleaved ectodomain of the paramyxovirus (hPIV3) fusion protein
Yin, H.S., Paterson, R.G., Wen, X., Lamb, R.A., Jardetzky, T.S.(2005) Proc Natl Acad Sci U S A 102: 9288-9293
- PubMed: 15964978
- DOI: https://doi.org/10.1073/pnas.0503989102
- Primary Citation of Related Structures:
1ZTM - PubMed Abstract:
Class I viral fusion proteins share common mechanistic and structural features but little sequence similarity. Structural insights into the protein conformational changes associated with membrane fusion are based largely on studies of the influenza virus hemagglutinin in pre- and postfusion conformations. Here, we present the crystal structure of the secreted, uncleaved ectodomain of the paramyxovirus, human parainfluenza virus 3 fusion (F) protein, a member of the class I viral fusion protein group. The secreted human parainfluenza virus 3 F forms a trimer with distinct head, neck, and stalk regions. Unexpectedly, the structure reveals a six-helix bundle associated with the postfusion form of F, suggesting that the anchor-minus ectodomain adopts a conformation largely similar to the postfusion state. The transmembrane anchor domains of F may therefore profoundly influence the folding energetics that establish and maintain a metastable, prefusion state.
Organizational Affiliation:
Department of Biochemistry, Molecular Biology, and Cell Biology, Howard Hughes Medical Institute, Northwestern University, Evanston, IL 60208-3500, USA.