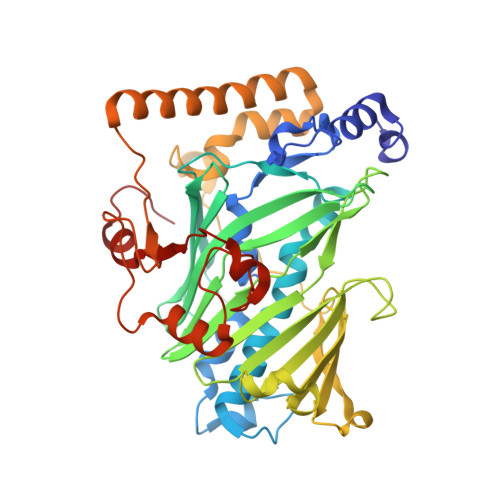
Structural mechanism for sterol sensing and transport by OSBP-related proteins
Im, Y.J., Raychaudhuri, S., Prinz, W.A., Hurley, J.H.(2005) Nature 437: 154-158
- PubMed: 16136145
- DOI: https://doi.org/10.1038/nature03923
- Primary Citation of Related Structures:
1ZHT, 1ZHW, 1ZHX, 1ZHY, 1ZHZ, 1ZI7 - PubMed Abstract:
The oxysterol-binding-protein (OSBP)-related proteins (ORPs) are conserved from yeast to humans, and are implicated in the regulation of sterol homeostasis and in signal transduction pathways. Here we report the structure of the full-length yeast ORP Osh4 (also known as Kes1) at 1.5-1.9 A resolution in complexes with ergosterol, cholesterol, and 7-, 20- and 25-hydroxycholesterol. We find that a single sterol molecule binds within a hydrophobic tunnel in a manner consistent with a transport function for ORPs. The entrance is blocked by a flexible amino-terminal lid and surrounded by basic residues that are critical for Osh4 function. The structure of the open state of a lid-truncated form of Osh4 was determined at 2.5 A resolution. Structural analysis and limited proteolysis show that sterol binding closes the lid and stabilizes a conformation favouring transport across aqueous barriers and signal transmission. The structure of Osh4 in the absence of ligand exposes potential phospholipid-binding sites that are positioned for membrane docking and sterol exchange. On the basis of these observations, we propose a model in which sterol and membrane binding promote reciprocal conformational changes that facilitate a sterol transfer and signalling cycle.
Organizational Affiliation:
Laboratory of Molecular Biology, National Institutes of Health, US Department of Health and Human Services, Bethesda, Maryland 20892, USA.