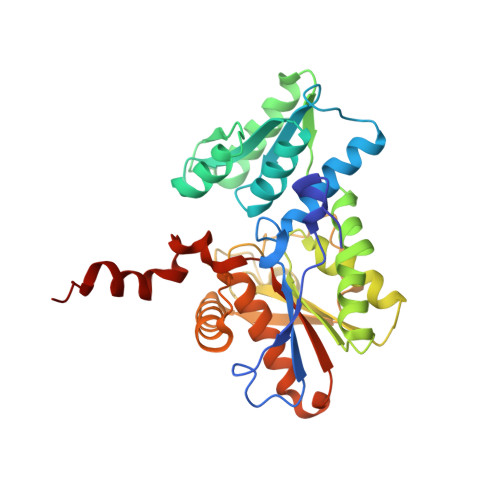
Molecular Basis of Cysteine Biosynthesis in Plants: STRUCTURAL AND FUNCTIONAL ANALYSIS OF O-ACETYLSERINE SULFHYDRYLASE FROM ARABIDOPSIS THALIANA.
Bonner, E.R., Cahoon, R.E., Knapke, S.M., Jez, J.M.(2005) J Biol Chem 280: 38803-38813
- PubMed: 16166087
- DOI: https://doi.org/10.1074/jbc.M505313200
- Primary Citation of Related Structures:
1Z7W, 1Z7Y - PubMed Abstract:
In plants, cysteine biosynthesis plays a central role in fixing inorganic sulfur from the environment and provides the only metabolic sulfide donor for the generation of methionine, glutathione, phytochelatins, iron-sulfur clusters, vitamin cofactors, and multiple secondary metabolites. O-Acetylserine sulfhydrylase (OASS) catalyzes the final step of cysteine biosynthesis, the pyridoxal 5'-phosphate (PLP)-dependent conversion of O-acetylserine into cysteine. Here we describe the 2.2 A resolution crystal structure of OASS from Arabidopsis thaliana (AtOASS) and the 2.7 A resolution structure of the AtOASS K46A mutant with PLP and methionine covalently linked as an external aldimine in the active site. Although the plant and bacterial OASS share a conserved set of amino acids for PLP binding, the structure of AtOASS reveals a difference from the bacterial enzyme in the positioning of an active site loop formed by residues 74-78 when methionine is bound. Site-directed mutagenesis, kinetic analysis, and ligand binding titrations probed the functional roles of active site residues. These experiments indicate that Asn(77) and Gln(147) are key amino acids for O-acetylserine binding and that Thr(74) and Ser(75) are involved in sulfur incorporation into cysteine. In addition, examination of the AtOASS structure and nearly 300 plant and bacterial OASS sequences suggest that the highly conserved beta8A-beta9A surface loop may be important for interaction with serine acetyltransferase, the other enzyme in cysteine biosynthesis. Initial protein-protein interaction experiments using AtOASS mutants targeted to this loop support this hypothesis.
Organizational Affiliation:
Donald Danforth Plant Science Center, St. Louis, Missouri 63132, USA.