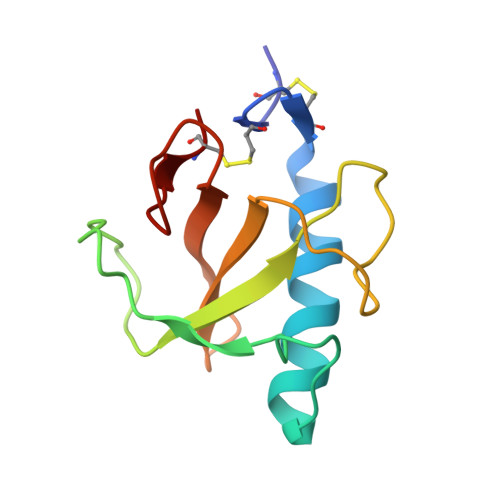
Purine activity of RNase T1RV is further improved by substitution of Trp59 by tyrosine
Czaja, R., Perbandt, M., Betzel, C., Hahn, U.(2005) Biochem Biophys Res Commun 336: 882-889
- PubMed: 16157302
- DOI: https://doi.org/10.1016/j.bbrc.2005.08.188
- Primary Citation of Related Structures:
1TTO - PubMed Abstract:
Ribonuclease T1 is an enzyme that cleaves single-stranded RNA with high specificity after guanylyl residues. Although this enzyme is a very good characterized protein with respect to structure and enzymatic function, we were only recently successful in generating RNase T1-RV, a variant where the specificity was changed from guanine to purine. As this change of substrate specificity was made at the cost of activity, the aim was now to further improve the overall activity of the enzyme. Therefore, we have substituted the tryptophan in position 59 by tyrosine. This substitution led to an increase of enzymatic activity in comparison to variant RV to 425%. As the extent of this enhancement is unique so far we have crystallized and analyzed the structure of this variant in order to get more insights into the reasons for this. Here, we present the crystal structure of this so-called RNase T1-R2 at 2.1A resolution. The structure was determined by molecular replacement using the coordinates of the RV variant (PDB entry: 1Q9E). The data were refined to an R-factor of 18.7% and R(free) of 24%, respectively. The asymmetric unit contains three molecules and the crystal packing is very similar to that of variant RV.
Organizational Affiliation:
Department of Chemistry, Division of Biochemistry and Molecular Biology, University of Hamburg, Martin-Luther-King-Platz 6, 20146 Hamburg, Germany.