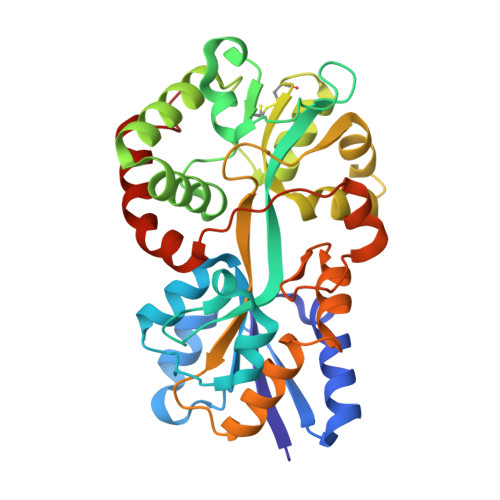
Structural basis for iron binding and release by a novel class of periplasmic iron-binding proteins found in gram-negative pathogens.
Shouldice, S.R., Skene, R.J., Dougan, D.R., Snell, G., McRee, D.E., Schryvers, A.B., Tari, L.W.(2004) J Bacteriol 186: 3903-3910
- PubMed: 15175304
- DOI: https://doi.org/10.1128/JB.186.12.3903-3910.2004
- Primary Citation of Related Structures:
1SI0, 1SI1 - PubMed Abstract:
We have determined the 1.35- and 1.45-A structures, respectively, of closed and open iron-loaded forms of Mannheimia haemolytica ferric ion-binding protein A. M. haemolytica is the causative agent in the economically important and fatal disease of cattle termed shipping fever. The periplasmic iron-binding protein of this gram-negative bacterium, which has homologous counterparts in many other pathogenic species, performs a key role in iron acquisition from mammalian host serum iron transport proteins and is essential for the survival of the pathogen within the host. The ferric (Fe(3+)) ion in the closed structure is bound by a novel asymmetric constellation of four ligands, including a synergistic carbonate anion. The open structure is ligated by three tyrosyl residues and a dynamically disordered solvent-exposed anion. Our results clearly implicate the synergistic anion as the primary mediator of global protein conformation and provide detailed insights into the molecular mechanisms of iron binding and release in the periplasm.
Organizational Affiliation:
Department of Microbiology and Infectious Diseases, University of Calgary, Calgary, AB, Canada.