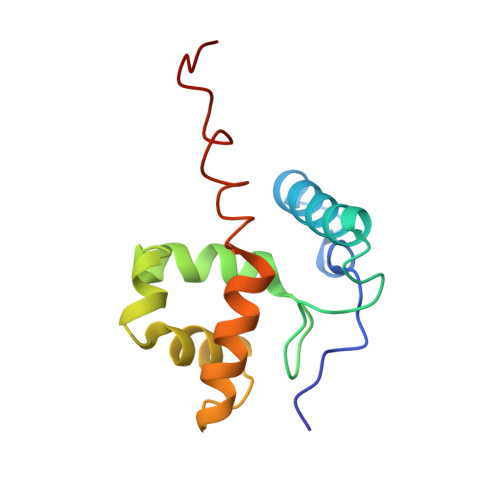
Structure and DNA-binding sites of the SWI1 AT-rich interaction domain (ARID) suggest determinants for sequence-specific DNA recognition.
Kim, S., Zhang, Z., Upchurch, S., Isern, N., Chen, Y.(2004) J Biol Chem 279: 16670-16676
- PubMed: 14722072
- DOI: https://doi.org/10.1074/jbc.M312115200
- Primary Citation of Related Structures:
1RYU - PubMed Abstract:
ARID (AT-rich interaction domain) is a homologous family of DNA-binding domains that occur in DNA-binding proteins from a wide variety of species, ranging from yeast to nematodes, insects, mammals, and plants. SWI1, a member of the SWI/SNF protein complex that is involved in chromatin remodeling during transcription, contains the ARID motif. The ARID domain of human SWI1 (also known as p270) does not select for a specific DNA sequence from a random sequence pool. The lack of sequence specificity shown by the SWI1 ARID domain stands in contrast to the other characterized ARID domains, which recognize specific AT-rich sequences. We have solved the three-dimensional structure of human SWI1 ARID using solution NMR methods. In addition, we have characterized nonspecific DNA binding by the SWI1 ARID domain. Results from this study indicate that a flexible, long, internal loop in the ARID motif is likely to be important for sequence-specific DNA recognition. The structure of the human SWI1 ARID domain also represents a distinct structural subfamily. Studies of ARID indicate that the boundary of DNA binding structural and functional domains can extend beyond the sequence homologous region in a homologous family of proteins. Structural studies of homologous domains such as the ARID family of DNA-binding domains should provide information to better predict the boundary of structural and functional domains in structural genomic studies.
Organizational Affiliation:
Division of Immunology, Beckman Research Institute of the City of Hope, Duarte, California 91010, USA.