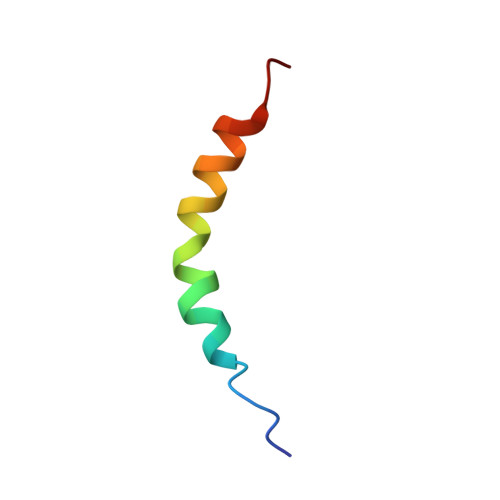
Structure and function of the membrane anchor domain of hepatitis C virus nonstructural protein 5A.
Penin, F., Brass, V., Appel, N., Ramboarina, S., Montserret, R., Ficheux, D., Blum, H.E., Bartenschlager, R., Moradpour, D.(2004) J Biol Chem 279: 40835-40843
- PubMed: 15247283
- DOI: https://doi.org/10.1074/jbc.M404761200
- Primary Citation of Related Structures:
1R7C, 1R7D, 1R7E, 1R7F, 1R7G - PubMed Abstract:
Hepatitis C virus (HCV) nonstructural protein 5A (NS5A) is a membrane-associated, essential component of the viral replication complex. Here, we report the three-dimensional structure of the membrane anchor domain of NS5A as determined by NMR spectroscopy. An alpha-helix extending from amino acid residue 5 to 25 was observed in the presence of different membrane mimetic media. This helix exhibited a hydrophobic, Trprich side embedded in detergent micelles, while the polar, charged side was exposed to the solvent. Thus, the NS5A membrane anchor domain forms an in-plane amphipathic alpha-helix embedded in the cytosolic leaflet of the membrane bilayer. Interestingly, mutations affecting the positioning of fully conserved residues located at the cytosolic surface of the helix impaired HCV RNA replication without interfering with the membrane association of NS5A. In conclusion, the NS5A membrane anchor domain constitutes a unique platform that is likely involved in specific interactions essential for the assembly of the HCV replication complex and that may represent a novel target for antiviral intervention.
Organizational Affiliation:
Institut de Biologie et Chimie des Protéines, CNRS-UMR 5086, IFR128 BioSciences, Lyon-Gerland, Lyon F-69367, Cedex 07, France.