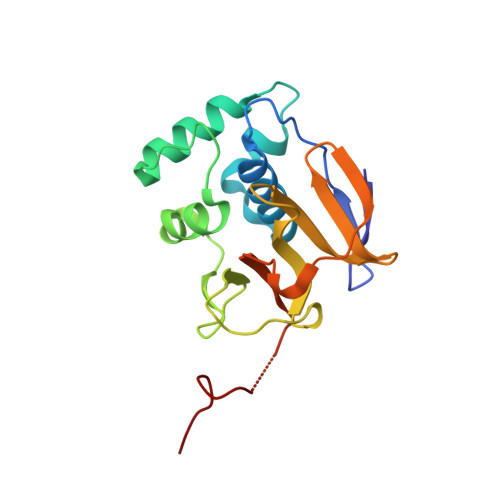
Structure of the Foot-and-Mouth Disease Virus Leader Protease: A Papain-Like Fold Adapted for Self-Processing and Eif4G Recognition.
Guarne, A., Tormo, J., Kirchweger, R., Pfistermueller, D., Fita, I., Skern, T.(1998) EMBO J 17: 7469
- PubMed: 9857201
- DOI: https://doi.org/10.1093/emboj/17.24.7469
- Primary Citation of Related Structures:
1QOL - PubMed Abstract:
The leader protease of foot-and-mouth disease virus, as well as cleaving itself from the nascent viral polyprotein, disables host cell protein synthesis by specific proteolysis of a cellular protein: the eukaryotic initiation factor 4G (eIF4G). The crystal structure of the leader protease presented here comprises a globular catalytic domain reminiscent of that of cysteine proteases of the papain superfamily, and a flexible C-terminal extension found intruding into the substrate-binding site of an adjacent molecule. Nevertheless, the relative disposition of this extension and the globular domain to each other supports intramolecular self-processing. The different sequences of the two substrates cleaved during viral replication, the viral polyprotein (at LysLeuLys/GlyAlaGly) and eIF4G (at AsnLeuGly/ArgThrThr), appear to be recognized by distinct features in a narrow, negatively charged groove traversing the active centre. The structure illustrates how the prototype papain fold has been adapted to the requirements of an RNA virus. Thus, the protein scaffold has been reduced to a minimum core domain, with the active site being modified to increase specificity. Furthermore, surface features have been developed which enable C-terminal self-processing from the viral polyprotein.
Organizational Affiliation:
Centre d'Investigació i Desenvolupament (CSIC), Jordi Girona Salgado 18-26, E-08034 Barcelona, Spain.